DECODERS platform: decoding locus-specific molecular interaction networks
Supported by the Transverse program of the LabEx (Laboratory of Excellence) Who Am I?, the DECODERS platform (established in November 2021) aims to develop and propose innovative proteomic and transcriptomic approaches within a consortium of laboratories gathered around a common scientific question: how do loci-specific molecular interactions networks participate in cellular identity and plasticity?
In this context, the DECODERS project aims to combine in situ proximity biotinylation and loci-specific targeting techniques to identify the molecular components (proteins or RNAs) of individual genomic regions, through mass spectrometry or high-throughput sequencing.
The projects includes scientific monitoring and methodological developments to recommend and make available the latest proximity biotinylation technologies.
Keywords: In situ Proximity-dependent Biotinylation (BioID), APEX2/TurboID, Proteomic, Transcriptomic, CRISPR/dCas9, HYPRO, AMAPEX


decoders (at) ijm.fr Bâtiment Buffon, 3rd floor, room 354B
Development and production of tools for molecular targeting and in situ biotinylation.
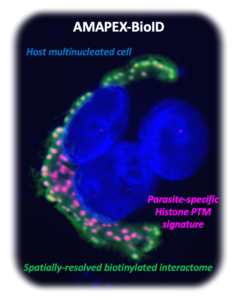
We use biotin ligase and peroxidase enzymes (TurboID or APEX2, respectively) targeted to individual loci to specifically label proximal interactants through in situ biotinylation. Targeting tools notably include CRISPR/dCas9 (CASPEX strategy), antibodies (AMAPEX strategy) or DNA probes (HYPRO strategy).
Design and implementation of experimental strategies
The DECODERS platform aims to support and assist with the design of appropriate experimental strategies for different research hypotheses and models. It implements biochemical and molecular approaches in close collaboration with the different teams and trains users in the experimental workflow.
High throughput data analysis
Mass spectrometry identification and differential quantitative analysis of interactants are performed in collaboration with the Proteomics platform of Institut Jacques Monod (ProteoSeine, Université Paris Cité).
Our services
– Collaboration with research teams for the design of proximity biotinylation projects on genomic regions.
– Realization of experimental procedures in molecular biology and biochemistry.
– Differential quantitative analysis of mass spectrometry data (in collaboration with ProteoSeine).
Scientific supervisor:
Benoit Palancade – Research Director, CNRS
Resources
The platform provides all the molecular and biochemical tools required to implement the different experimental approaches.
– Sharable database of in situ biotinylation protocols.
– Plasmid library (APEX2, TurboID, Split TurboID, dCas9…) and cloning tools.
– Recombinant proteins (APEX2-protein A; APEX2-protein A/G; APEX2-HYPRO).
– Biotinylation and biotin enrichment reagents.
– Digoxigenin labeling kits for DNA probes (HYPRO strategy).
– Antibodies for validation by immunofluorescence.
We are working with research groups involved in the Transverse program of the WhoAmI? laboratory of excellence (LabEx) supported by Université Paris Cité:
UMR7592, Institut Jacques Monod
Jean Charles Cadoret
Sandra Duharcourt
Maxim Greenberg
Benoit Palancade
Marie Noëlle Prioleau
Vanessa Ribes
ProtéoSeine facility
UMR7216, Epigenetic and cell fate
Pierre Antoine Defossez
Claire Francastel
Valérie Mezger
Claire Rougeulle
Sophie Polo
Jonathan Weitzman
GENIE (Genomic engineering in epigenetics) facility
Institut Cochin
Julie Chaumeil
Stéphane Emiliani
Bernard Lopez
Pascal Maire
Daniel Vaiman

