Research
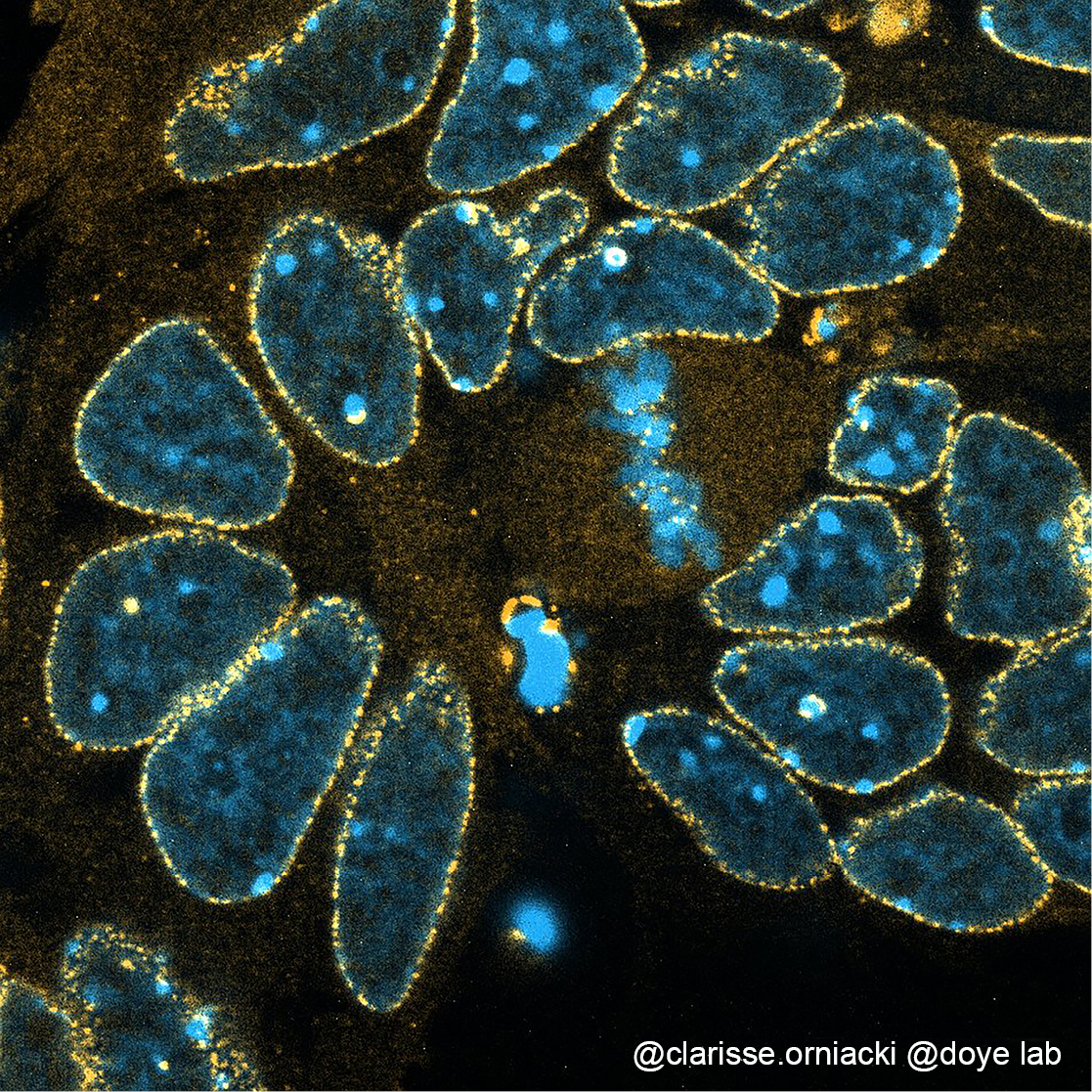
Nuclear Pore Complexes (NPCs) are elaborate structures embedded in the nuclear envelope and composed of multiple copies of about 30 different proteins termed nucleoporins (Nups). Our previous work contributed to the characterization of a major structural sub-complex of NPCs, the Nup107-160 complex (also called “Y-complex“, Figure 1), and to highlight its role in NPC assembly, its contribution to various stages of mitotic progression, and its involvement in embryonic stem cell differentiation.
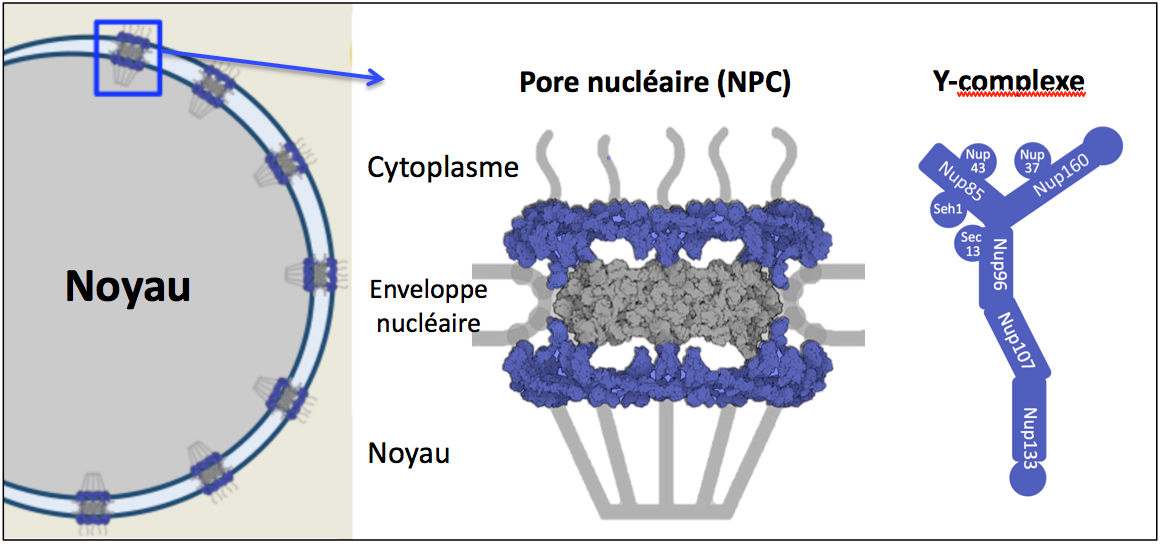
Figure 1: Schematic representation of a nuclear pore complex and the components of the Y-complex.
We are currently pursuing the study of the functions, regulation and direct partners of these nucleoporins in these “unconventional” activities. To do so, we use imaging, biochemical and genetic approaches (CRISPR/Cas9 strategies), mainly in pluripotent stem cells (mESCs and hiPSCs) and their 2D and 3D-differentiated derivatives.
Projet 1: Implication of Y-complex Nups in mouse embryonic stem cell (mESC) differentiation
Our team has previously shown that several Y-complex Nups, namely Nup133, Seh1 and Nup43, are dispensable for cell viability in pluripotent mouse embryonic stem cells (mESCs) but required for their survival during differentiation (Lupu et al., 2008; Gonzalez-Estevez, Verrico et al., 2021). In addition, we demonstrated that in pluripotent mESCs, Nup133 is necessary for proper assembly of the nuclear pore basket (Souquet et al., 2018) whereas mutations affecting the short arm of the Y-complex (Nup85, Seh1 and Nup43) negatively impact NPC density (Gonzalez-Estevez, Verrico et al., 2021).
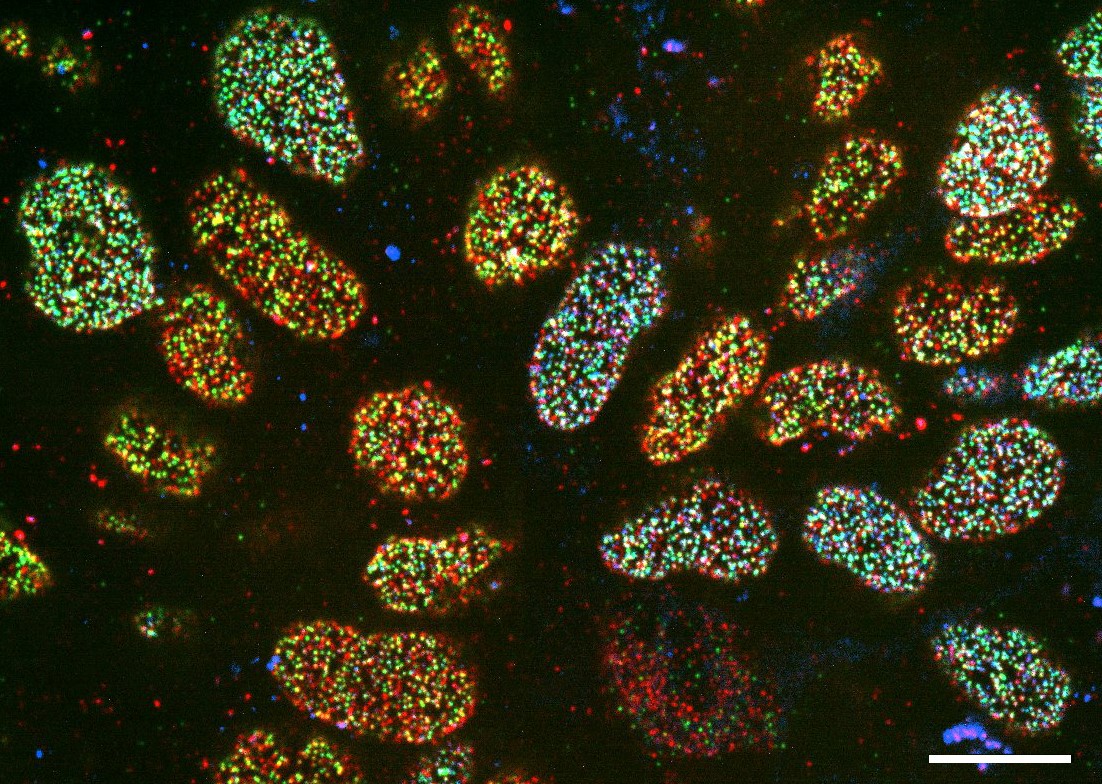
Figure 2: Nuclear pore complexes in neuronal progenitors derived from mESCs expressing various levels of GFP-Nup133 (represented in blue), and labeled with antibodies directed against Nup153 (red) and TPR (green). Scale bar, 10 µm.
Additionally, an RNA-seq analysis enabled the identification of potential target genes regulated by Nup133 during early steps of neuronal differentiation (unpublished results from our team, collaboration with the Genomics facility from IBENS, Paris, France). We are currently validating these target genes and characterizing the mechanisms by which the Y-complex contributes to their regulation.
Project 2: Molecular mechanisms leading to microcephaly and/or hereditary nephrotic syndromes upon nucleoporin defects
In recent years, several teams have identified mutations in genes encoding structural Nups, notably Y-complex Nups, in patients with an early-onset and severe kidney disease (steroid-resistant nephrotic syndrome, SRNS), and/or neurodevelopmental disorders, mainly microcephaly. However, the reason why mutations in these ubiquitously expressed Nups lead to pathologies mainly affecting the kidney and/or the brain remains a mystery.
To mimic the ”disease” condition associated with distinct nucleoporin mutations, notably those causing SRNS and/or microcephaly, we are using human induced pluripotent stem cells (hiPSCs) generated either from peripheral blood cells from patients with NUP mutations (collaboration with C. Antignac and the iPSCs facility from Imagine Institute, Paris) or from healthy individuals in which mutations are introduced by CRISPR/Cas9-mediated gene editing (with help of the enSCORE plateform – labex who am I?).
To investigate organ- and cell type-specific and thus pathologically relevant phenotypes, the iPSCs are differentiated towards 3D brain organoids (Figure 3). (with help from the enSCORE plateform) or kidney organoids (in collaboration with C. Antignac and G. Mollet, Imagine Institute).
In both pluripotent cells and their derived organoids, we then study the impact of Y-Nups mutations on NPC assembly, nuclear transport, cell division and migration, and gene regulation. We also aim to characterize Y-Nups partners that may explain the tissue-specific alterations observed in patients.
By elucidating the role of these nucleoporins, our work should contribute to understanding the mechanisms by which mutations or misregulation of these nucleoporins lead to various pathologies.
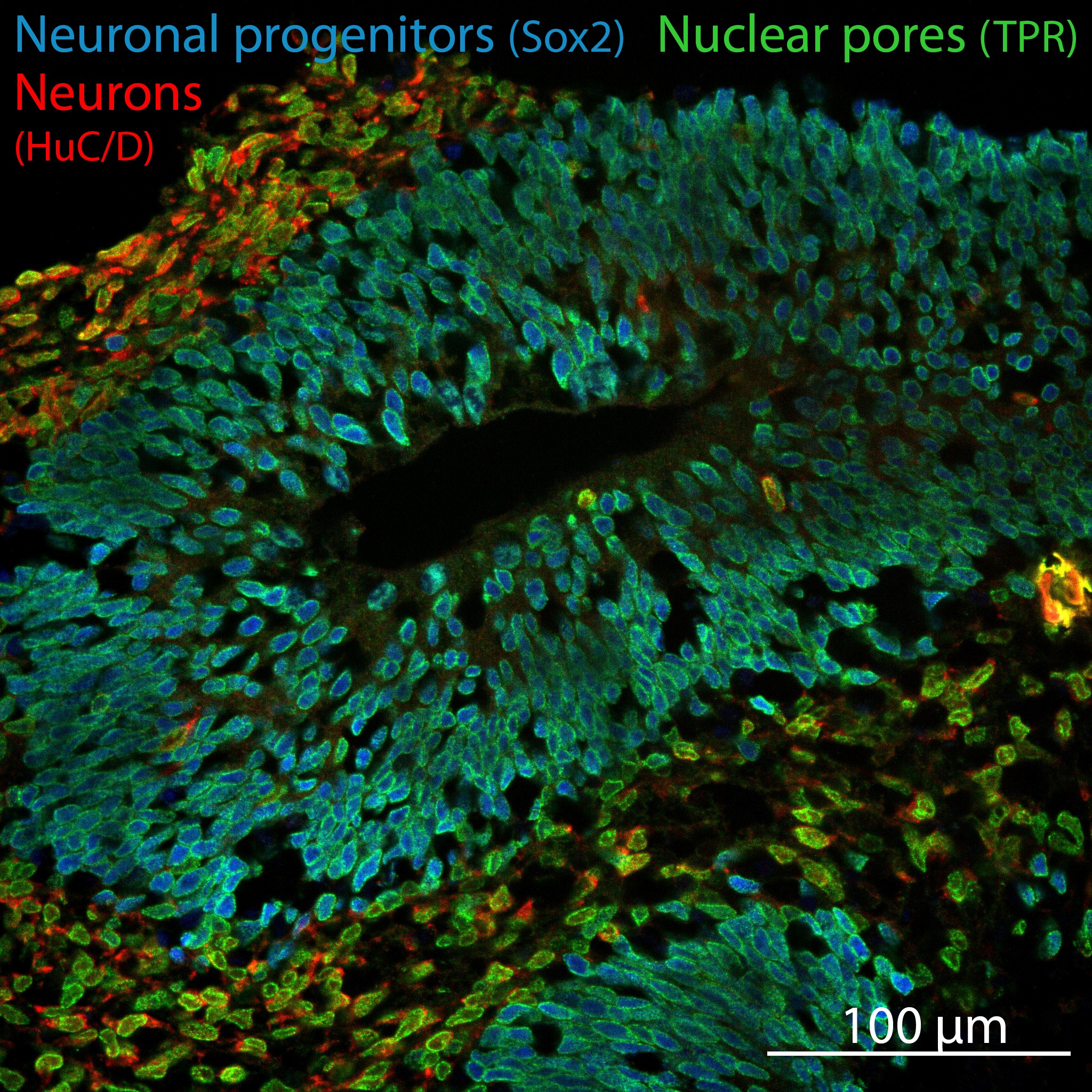
Figure 3: Cryosection of a cerebral organoids at day 38 of differentiation.
Models: pluripotent stem cells (mESCs and hiPSCs), organoids, primary and cancer cell lines
Methods: genome editing, cell differentiation, cell imaging, transcriptomics, biochemistry, yeast-two-hybrid
Members
Team leader
Valerie DOYE,
Researcher,
DOYE LAB+33 (0)1 57 27 81 39, room RB08B
Members
Brigitte BUENDIA,
Researcher,
DOYE LAB+33 (0)1 57 27 80 60, room 344B
Marie DIDELOT,
Intern,
DOYE LAB+33 (0)1 57 27 80 62, room 314B
Roger KARESS,
Emeritus researcher,
DOYE LAB+33 (0)1 57 27 80 95, room 344B
Fernanda LOPEZ GARCIA,
Intern,
DOYE LAB Stephanie MORCHOISNE,
Biology engineer,
DOYE LAB+33 (0)1 57 27 80 61, room 342B
Gulin OZEK,
Intern,
DOYE LAB+33 (0)1 57 27 80 62, room 314B
Ona PRAT PLANAS,
PhD student,
DOYE LAB+33 (0)1 57 27 80 62, room 342B
Nathalie VADROT,
Biology engineer,
DOYE LAB+33 (0)1 57 27 80 61, room 342B
To contact a member of the team by e-mail: name.surname@ijm.fr
Selected publications
Nuclear pores safeguard the integrity of the nuclear envelope. Taniguchi, R., Orniacki, C., Kreysing, J. P., Zila, V., Zimmerli, C. E., Böhm, S., Turoňová, B., Kräusslich, H.-G., Doye, V., & Beck, M. (2025). Nature Cell Biology, 1–14. https://doi.org/10.1038/s41556-025-01648-3
Y-complex nucleoporins independently contribute to nuclear pore assembly and gene regulation in neuronal progenitors. Orniacki C., Verrico A., Pelletier S., Souquet B., Coulpier F., Jourdren L., Benetti S., & Doye V. J Cell Sci. 2023 Jun 1;136(11):jcs261151. doi: 10.1242/jcs.261151. PMID: 37305998 https://doi.org/10.1242/jcs.261151
Moderate Nucleoporin 133 deficiency leads to glomerular damage in zebrafish. Cianciolo Cosentino C, Berto A, Pelletier S, Hari M, Loffing J, Neuhauss SCF, Doye V. Sci Rep. 2019 Mar 18;9(1):4750. doi: 10.1038/s41598-019-41202-4. PMID: 30894603
Nup133 Is Required for Proper Nuclear Pore Basket Assembly and Dynamics in Embryonic Stem Cells. Souquet B, Freed E, Berto A, Andric V, Audugé N, Reina-San-Martin B, Lacy E, Doye V. Cell Rep. 2018 May 22;23(8):2443-2454. doi: 10.1016/j.celrep.2018.04.070. PMID: 29791854
Disentangling the molecular determinants for Cenp-F localization to nuclear pores and kinetochores. Berto A, Yu J, Morchoisne-Bolhy S, Bertipaglia C, Vallee R, Dumont J, Ochsenbein F, Guerois R, Doye V. EMBO Rep. 2018 May;19(5):e44742. doi: 10.15252/embr.201744742. PMID: 29632243
Dynein recruitment to nuclear pores activates apical nuclear migration and mitotic entry in brain progenitor cells. Hu DJ, Baffet AD, Nayak T, Akhmanova A, Doye V, Vallee RB. Cell. 2013 Sep 12;154(6):1300-13. doi: 10.1016/j.cell.2013.08.024. PMID: 24034252
A Nup133-dependent NPC-anchored network tethers centrosomes to the nuclear envelope in prophase. Bolhy S, Bouhlel I, Dultz E, Nayak T, Zuccolo M, Gatti X, Vallee R, Ellenberg J, Doye V. J Cell Biol. 2011 Mar 7;192(5):855-71. doi: 10.1083/jcb.201007118. PMID: 21383080
Nuclear pore composition regulates neural stem/progenitor cell differentiation in the mouse embryo. Lupu F, Alves A, Anderson K, Doye V, Lacy E. Dev Cell. 2008 Jun;14(6):831-42. doi: 10.1016/j.devcel.2008.03.011. PMID: 18539113
The human Nup107-160 nuclear pore subcomplex contributes to proper kinetochore functions. Zuccolo M, Alves A, Galy V, Bolhy S, Formstecher E, Racine V, Sibarita JB, Fukagawa T, Shiekhattar R, Yen T, Doye V. EMBO J. 2007 Apr 4;26(7):1853-64. doi: 10.1038/sj.emboj.7601642. PMID: 17363900
The conserved Nup107-160 complex is critical for nuclear pore complex assembly. Walther TC, Alves A, Pickersgill H, Loïodice I, Hetzer M, Galy V, Hülsmann BB, Köcher T, Wilm M, Allen T, Mattaj IW, Doye V. Cell. 2003 Apr 18;113(2):195-206. doi: 10.1016/s0092-8674(03)00235-6. PMID: 12705868
Publications
Publications
2913254
YZUD6P7F
1
apa
50
date
desc
8305
https://www.ijm.fr/wp-content/plugins/zotpress/
%7B%22status%22%3A%22success%22%2C%22updateneeded%22%3Afalse%2C%22instance%22%3Afalse%2C%22meta%22%3A%7B%22request_last%22%3A0%2C%22request_next%22%3A0%2C%22used_cache%22%3Atrue%7D%2C%22data%22%3A%5B%7B%22key%22%3A%22IBEIZ7FA%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Vadrot%20et%20al.%22%2C%22parsedDate%22%3A%222026-01-21%22%2C%22numChildren%22%3A1%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BVadrot%2C%20N.%2C%20Moulin%2C%20M.%2C%20Ferreiro%2C%20A.%2C%20Richard%2C%20P.%2C%20%26amp%3B%20Buendia%2C%20B.%20%282026%29.%20LAP2%20Isoform%20Profile%20in%20Heart%20Ageing%20and%20in%20Cardiac%20Cell%20Proliferation%20and%20Differentiation%3A%20Input%20From%20CRISPR-Cas9-mediated%20LAP2a%20Knockdown%20in%20H9C2.%20%26lt%3Bi%26gt%3BInternational%20Journal%20of%20Medical%20Sciences%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B23%26lt%3B%5C%2Fi%26gt%3B%283%29%2C%20741%26%23×2013%3B757.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.7150%5C%2Fijms.114095%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.7150%5C%2Fijms.114095%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22LAP2%20Isoform%20Profile%20in%20Heart%20Ageing%20and%20in%20Cardiac%20Cell%20Proliferation%20and%20Differentiation%3A%20Input%20From%20CRISPR-Cas9-mediated%20LAP2a%20Knockdown%20in%20H9C2%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Nathalie%22%2C%22lastName%22%3A%22Vadrot%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Maryline%22%2C%22lastName%22%3A%22Moulin%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Ana%22%2C%22lastName%22%3A%22Ferreiro%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Pascale%22%2C%22lastName%22%3A%22Richard%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Brigitte%22%2C%22lastName%22%3A%22Buendia%22%7D%5D%2C%22abstractNote%22%3A%22%22%2C%22date%22%3A%222026%5C%2F1%5C%2F21%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.7150%5C%2Fijms.114095%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22https%3A%5C%2F%5C%2Fwww.medsci.org%5C%2Fv23p0741.htm%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221449-1907%22%2C%22language%22%3A%22en%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222026-03-10T10%3A02%3A12Z%22%7D%7D%2C%7B%22key%22%3A%22D2N4HSDA%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Taniguchi%20et%20al.%22%2C%22parsedDate%22%3A%222025-04-09%22%2C%22numChildren%22%3A1%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BTaniguchi%2C%20R.%2C%20Orniacki%2C%20C.%2C%20Kreysing%2C%20J.%20P.%2C%20Zila%2C%20V.%2C%20Zimmerli%2C%20C.%20E.%2C%20B%26%23xF6%3Bhm%2C%20S.%2C%20Turo%26%23×148%3Bov%26%23xE1%3B%2C%20B.%2C%20Kr%26%23xE4%3Busslich%2C%20H.-G.%2C%20Doye%2C%20V.%2C%20%26amp%3B%20Beck%2C%20M.%20%282025%29.%20Nuclear%20pores%20safeguard%20the%20integrity%20of%20the%20nuclear%20envelope.%20%26lt%3Bi%26gt%3BNature%20Cell%20Biology%26lt%3B%5C%2Fi%26gt%3B%2C%201%26%23×2013%3B14.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41556-025-01648-3%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41556-025-01648-3%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Nuclear%20pores%20safeguard%20the%20integrity%20of%20the%20nuclear%20envelope%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Reiya%22%2C%22lastName%22%3A%22Taniguchi%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Clarisse%22%2C%22lastName%22%3A%22Orniacki%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Jan%20Philipp%22%2C%22lastName%22%3A%22Kreysing%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Vojtech%22%2C%22lastName%22%3A%22Zila%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Christian%20E.%22%2C%22lastName%22%3A%22Zimmerli%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Stefanie%22%2C%22lastName%22%3A%22B%5Cu00f6hm%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Beata%22%2C%22lastName%22%3A%22Turo%5Cu0148ov%5Cu00e1%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Hans-Georg%22%2C%22lastName%22%3A%22Kr%5Cu00e4usslich%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Martin%22%2C%22lastName%22%3A%22Beck%22%7D%5D%2C%22abstractNote%22%3A%22Nuclear%20pore%20complexes%20%28NPCs%29%20mediate%20nucleocytoplasmic%20exchange%2C%20which%20is%20essential%20for%20eukaryotes.%20Mutations%20in%20the%20central%20scaffolding%20components%20of%20NPCs%20are%20associated%20with%20genetic%20diseases%2C%20but%20how%20they%20manifest%20only%20in%20specific%20tissues%20remains%20unclear.%20This%20is%20exemplified%20in%20Nup133-deficient%20mouse%20embryonic%20stem%20cells%2C%20which%20grow%20normally%20during%20pluripotency%2C%20but%20differentiate%20poorly%20into%20neurons.%20Here%2C%20using%20an%20innovative%20in%20situ%20structural%20biology%20approach%2C%20we%20show%20that%20Nup133%5Cu2212%5C%2F%5Cu2212%20mouse%20embryonic%20stem%20cells%20have%20heterogeneous%20NPCs%20with%20non-canonical%20symmetries%20and%20missing%20subunits.%20During%20neuronal%20differentiation%2C%20Nup133-deficient%20NPCs%20frequently%20disintegrate%2C%20resulting%20in%20abnormally%20large%20nuclear%20envelope%20openings.%20We%20propose%20that%20the%20elasticity%20of%20the%20NPC%20scaffold%20has%20a%20protective%20function%20for%20the%20nuclear%20envelope%20and%20that%20its%20perturbation%20becomes%20critical%20under%20conditions%20that%20impose%20an%20increased%20mechanical%20load%20onto%20nuclei.%22%2C%22date%22%3A%222025-04-09%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1038%5C%2Fs41556-025-01648-3%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22https%3A%5C%2F%5C%2Fwww.nature.com%5C%2Farticles%5C%2Fs41556-025-01648-3%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221476-4679%22%2C%22language%22%3A%22en%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222025-04-11T12%3A29%3A26Z%22%7D%7D%2C%7B%22key%22%3A%22FGBLJQ3Y%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Dultz%20and%20Doye%22%2C%22parsedDate%22%3A%222025-02-01%22%2C%22numChildren%22%3A1%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BDultz%2C%20E.%2C%20%26amp%3B%20Doye%2C%20V.%20%282025%29.%20Opening%20the%20gate%3A%20Complexity%20and%20modularity%20of%20the%20nuclear%20pore%20scaffold%20and%20basket.%20%26lt%3Bi%26gt%3BCurrent%20Opinion%20in%20Cell%20Biology%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B92%26lt%3B%5C%2Fi%26gt%3B%2C%20102461.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.ceb.2024.102461%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.ceb.2024.102461%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Opening%20the%20gate%3A%20Complexity%20and%20modularity%20of%20the%20nuclear%20pore%20scaffold%20and%20basket%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Elisa%22%2C%22lastName%22%3A%22Dultz%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22Nuclear%20pore%20complexes%20%28NPCs%29%20are%20giant%20molecular%20assemblies%20that%20form%20the%20gateway%20between%20the%20nucleus%20and%20the%20cytoplasm%20and%20accommodate%20the%20bidirectional%20transport%20of%20a%20large%20variety%20of%20cargoes.%20Recent%20years%20have%20seen%20tremendous%20advances%20in%20our%20understanding%20of%20their%20building%20principles%20and%20have%20in%20particular%20called%20attention%20to%20the%20flexibility%20and%20variability%20of%20NPC%20composition%20and%20structure.%20Here%2C%20we%20review%20these%20recent%20advances%20and%20discuss%20how%20the%20newest%20technologies%20push%20the%20boundaries%20of%20nuclear%20pore%20research%20forward%2C%20with%20a%20specific%20highlight%20on%20the%20NPC%20scaffold%20and%20a%20prominent%20pore%20appendage%2C%20the%20nuclear%20basket%2C%20whose%20architecture%20has%20long%20been%20elusive.%22%2C%22date%22%3A%222025-02-01%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1016%5C%2Fj.ceb.2024.102461%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22https%3A%5C%2F%5C%2Fwww.sciencedirect.com%5C%2Fscience%5C%2Farticle%5C%2Fpii%5C%2FS0955067424001406%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%220955-0674%22%2C%22language%22%3A%22%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222025-01-23T08%3A57%3A43Z%22%7D%7D%2C%7B%22key%22%3A%22G4GB3Z2I%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Nobari%20et%20al.%22%2C%22parsedDate%22%3A%222023-10-01%22%2C%22numChildren%22%3A2%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BNobari%2C%20P.%2C%20Doye%2C%20V.%2C%20%26amp%3B%20Boumendil%2C%20C.%20%282023%29.%20Metazoan%20nuclear%20pore%20complexes%20in%20gene%20regulation%20and%20genome%20stability.%20%26lt%3Bi%26gt%3BDNA%20Repair%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B130%26lt%3B%5C%2Fi%26gt%3B%2C%20103565.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.dnarep.2023.103565%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.dnarep.2023.103565%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Metazoan%20nuclear%20pore%20complexes%20in%20gene%20regulation%20and%20genome%20stability%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Parisa%22%2C%22lastName%22%3A%22Nobari%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Charlene%22%2C%22lastName%22%3A%22Boumendil%22%7D%5D%2C%22abstractNote%22%3A%22The%20nuclear%20pore%20complexes%20%28NPCs%29%2C%20one%20of%20the%20hallmarks%20of%20eukaryotic%20nuclei%2C%20allow%20selective%20transport%20of%20macromolecules%20between%20the%20cytoplasm%20and%20the%20nucleus.%20Besides%20this%20canonical%20function%2C%20an%20increasing%20number%20of%20additional%20roles%20have%20been%20attributed%20to%20the%20NPCs%20and%20their%20constituents%2C%20the%20nucleoporins.%20Here%20we%20review%20recent%20insights%20into%20the%20mechanisms%20by%20which%20NPCs%20and%20nucleoporins%20affect%20transcription%20and%20DNA%20repair%20in%20metazoans.%20In%20the%20first%20part%2C%20we%20discuss%20how%20gene%20expression%20can%20be%20affected%20by%20the%20localization%20of%20genome-nucleoporin%20interactions%20at%20pores%20or%20%5Cu201coff-pores%5Cu201d%2C%20by%20the%20role%20of%20nucleoporins%20in%20chromatin%20organization%20at%20different%20scales%2C%20or%20by%20the%20physical%20properties%20of%20nucleoporins.%20In%20the%20second%20part%2C%20we%20review%20the%20contribution%20of%20NPCs%20to%20genome%20stability%2C%20including%20transport-dependent%20and%20-independent%20functions%20and%20the%20role%20of%20positioning%20at%20NPCs%20in%20the%20repair%20of%20heterochromatic%20breaks%20and%20the%20regulation%20of%20replication%20stress.%22%2C%22date%22%3A%222023-10-01%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1016%5C%2Fj.dnarep.2023.103565%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22https%3A%5C%2F%5C%2Fwww.sciencedirect.com%5C%2Fscience%5C%2Farticle%5C%2Fpii%5C%2FS1568786423001192%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221568-7864%22%2C%22language%22%3A%22%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222023-09-12T08%3A20%3A11Z%22%7D%7D%2C%7B%22key%22%3A%22K25BXYEV%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Orniacki%20et%20al.%22%2C%22parsedDate%22%3A%222023-05-17%22%2C%22numChildren%22%3A2%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BOrniacki%2C%20C.%2C%20Verrico%2C%20A.%2C%20Pelletier%2C%20S.%2C%20Souquet%2C%20B.%2C%20Coulpier%2C%20F.%2C%20Jourdren%2C%20L.%2C%20Benetti%2C%20S.%2C%20%26amp%3B%20Doye%2C%20V.%20%282023%29.%20Y-complex%20nucleoporins%20independently%20contribute%20to%20nuclear%20pore%20assembly%20and%20gene%20regulation%20in%20neuronal%20progenitors.%20%26lt%3Bi%26gt%3BJournal%20of%20Cell%20Science%26lt%3B%5C%2Fi%26gt%3B%2C%20jcs.261151.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1242%5C%2Fjcs.261151%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1242%5C%2Fjcs.261151%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Y-complex%20nucleoporins%20independently%20contribute%20to%20nuclear%20pore%20assembly%20and%20gene%20regulation%20in%20neuronal%20progenitors%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Clarisse%22%2C%22lastName%22%3A%22Orniacki%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Annalisa%22%2C%22lastName%22%3A%22Verrico%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22St%5Cu00e9phane%22%2C%22lastName%22%3A%22Pelletier%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Benoit%22%2C%22lastName%22%3A%22Souquet%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Fanny%22%2C%22lastName%22%3A%22Coulpier%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Laurent%22%2C%22lastName%22%3A%22Jourdren%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Serena%22%2C%22lastName%22%3A%22Benetti%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22Besides%20assembling%20nuclear%20pore%20complexes%2C%20the%20conduits%20of%20nuclear%20transport%2C%20many%20nucleoporins%20also%20contribute%20to%20chromatin%20organization%20and%20gene%20expression%2C%20with%20critical%20roles%20in%20development%20and%20pathologies.%20We%20previously%20reported%20that%20Nup133%20and%20Seh1%2C%20two%20components%20of%20the%20Y-complex%20subassembly%20of%20the%20nuclear%20pore%20scaffold%2C%20are%20dispensable%20for%20mouse%20embryonic%20stem%20cell%20viability%20but%20required%20for%20their%20survival%20during%20neuroectodermal%20differentiation.%20Here%2C%20a%20transcriptomic%20analysis%20revealed%20that%20Nup133%20regulates%20a%20subset%20of%20genes%20at%20early%20stages%20of%20neuroectodermal%20differentiation%2C%20including%20Lhx1%20and%20Nup210L%2C%20encoding%20a%20newly%20validated%20nucleoporin.%20These%20genes%20are%20also%20misregulated%20in%20Nup133%5Cu25b5Mid%20neuronal%20progenitors%2C%20in%20which%20nuclear%20pore%20basket%20assembly%20is%20impaired.%20However%2C%20a%20four-fold%20reduction%20of%20Nup133%2C%20despite%20also%20affecting%20basket%20assembly%2C%20is%20not%20sufficient%20to%20alter%20Nup210L%20and%20Lhx1%20expression.%20Finally%2C%20these%20two%20genes%20are%20also%20misregulated%20in%20Seh1-deficient%20neural%20progenitors%20only%20showing%20a%20mild%20reduction%20in%20nuclear%20pore%20density.%20Together%20these%20data%20reveal%20a%20shared%20function%20of%20Y-complex%20nucleoporins%20in%20gene%20regulation%20during%20neuroectodermal%20differentiation%2C%20apparently%20independent%20of%20nuclear%20pore%20basket%20integrity.%22%2C%22date%22%3A%222023-05-17%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1242%5C%2Fjcs.261151%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221477-9137%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222023-05-23T10%3A21%3A47Z%22%7D%7D%2C%7B%22key%22%3A%22QR7AUZT7%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Bouglita%20et%20al.%22%2C%22parsedDate%22%3A%222022-09-26%22%2C%22numChildren%22%3A1%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BBouglita%2C%20W.%2C%20Rabhi%2C%20S.%2C%20Raich%2C%20N.%2C%20Bouabid%2C%20C.%2C%20Belghith%2C%20C.%2C%20Slimani%2C%20O.%2C%20Hkimi%2C%20C.%2C%20Ghedira%2C%20K.%2C%20Karess%2C%20R.%20E.%2C%20Guizani-Tabbane%2C%20L.%2C%20Attia%2C%20L.%2C%20%26amp%3B%20Rabhi%2C%20I.%20%282022%29.%20Microbiological%20and%20molecular%20screening%20of%20Candida%20spp.%20isolated%20from%20genital%20tract%20of%20asymptomatic%20pregnant%20women.%20%26lt%3Bi%26gt%3BJournal%20of%20Medical%20Microbiology%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B71%26lt%3B%5C%2Fi%26gt%3B%289%29%2C%20001589.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1099%5C%2Fjmm.0.001589%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1099%5C%2Fjmm.0.001589%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Microbiological%20and%20molecular%20screening%20of%20Candida%20spp.%20isolated%20from%20genital%20tract%20of%20asymptomatic%20pregnant%20women%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Wafa%22%2C%22lastName%22%3A%22Bouglita%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Sameh%22%2C%22lastName%22%3A%22Rabhi%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Natacha%22%2C%22lastName%22%3A%22Raich%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Cyrine%22%2C%22lastName%22%3A%22Bouabid%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Cyrine%22%2C%22lastName%22%3A%22Belghith%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Olfa%22%2C%22lastName%22%3A%22Slimani%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Chaima%22%2C%22lastName%22%3A%22Hkimi%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Kais%22%2C%22lastName%22%3A%22Ghedira%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Roger%20E.%22%2C%22lastName%22%3A%22Karess%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Lamia%22%2C%22lastName%22%3A%22Guizani-Tabbane%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Leila%22%2C%22lastName%22%3A%22Attia%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Imen%22%2C%22lastName%22%3A%22Rabhi%22%7D%5D%2C%22abstractNote%22%3A%22Introduction.%20Candida%20spp.%20may%20cause%20opportunistic%20infections%20called%20vulvovaginal%20candidiasis%20%28VVC%29%2C%20which%20is%20estimated%20to%20be%20the%20second%20most%20common%20cause%20of%20vaginitis%20worldwide.%20Gap%20Statement.%20Under%20various%20circumstances%2C%20VVC%20could%20compromise%20pregnancy%20outcomes.%20Emerging%20data%20suggests%20that%20VVC%20during%20pregnancy%20may%20be%20associated%20with%20increased%20risk%20of%20complications%20and%20congenital%20cutaneous%20candidiasis.%20Aim.%20To%20assess%20the%20prevalence%20of%20Candida%20spp.%20in%20asymptomatic%20pregnant%20women%20and%20determine%20the%20susceptibility%20of%20the%20isolates%20to%20antifungal%20drugs.%20Methodology.%20In%20a%20prospective%20cohort%2C%2065%20high%20vaginal%20swab%20samples%20of%20consented%20pregnant%20women.%20Candida%20isolates%20were%20identified%20using%20both%20microbiological%20and%20molecular%20tools%20and%20drug%20susceptibilities%20were%20profiled.%20Results.%20The%20prevalence%20of%20VVC%20among%20our%20study%20participants%20was%2037%20%25%2C%2024%20of%20the%2065%20asymptomatic%20pregnant%20women%20show%20Candida%20spp.%20colonization.%20C.%20albicans%20was%20the%20most%20common%20species%2061%20%25%2C%20followed%20by%20C.%20glabrata%2039%20%25.%20In%20addition%2C%20a%20significant%20fraction%20of%20the%20isolated%20colonies%20showed%20resistance%20to%20Fluconazole%2C%20with%20a%20ratio%20of%2063%20%25%20for%20C.%20albicans%20isolates%20and%2016%20%25%20for%20Candida%20glabrata%20isolates.%20Moreover%2C%20relative%20quantification%20of%20genes%20related%20to%20resistance%20to%20fluconazole%2C%20CDR1%2C%20ERG11%20as%20well%20as%20HWP1%2C%20showed%20a%20significant%20change%20compared%20to%20controls.%20Conclusion.%20Monitoring%20of%20vaginal%20Candida%20colonization%20before%20the%20third%20trimester%20of%20pregnancy%2C%20that%20could%20reduce%20congenital%20Candida%20colonization%20and%20risk%20of%20pregnancy%20complications.%22%2C%22date%22%3A%222022%5C%2F09%5C%2F26%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1099%5C%2Fjmm.0.001589%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22https%3A%5C%2F%5C%2Fwww.microbiologyresearch.org%5C%2Fcontent%5C%2Fjournal%5C%2Fjmm%5C%2F10.1099%5C%2Fjmm.0.001589%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%220022-2615%2C%201473-5644%22%2C%22language%22%3A%22en%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222023-05-23T12%3A07%3A18Z%22%7D%7D%2C%7B%22key%22%3A%22T89XWD3E%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Olley%20et%20al.%22%2C%22parsedDate%22%3A%222021-05-25%22%2C%22numChildren%22%3A3%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BOlley%2C%20G.%2C%20Pradeepa%2C%20M.%20M.%2C%20Grimes%2C%20G.%20R.%2C%20Piquet%2C%20S.%2C%20Polo%2C%20S.%20E.%2C%20FitzPatrick%2C%20D.%20R.%2C%20Bickmore%2C%20W.%20A.%2C%20%26amp%3B%20Boumendil%2C%20C.%20%282021%29.%20Cornelia%26%23xA0%3Bde%20Lange%20syndrome-associated%20mutations%20cause%20a%20DNA%20damage%20signalling%20and%20repair%20defect.%20%26lt%3Bi%26gt%3BNature%20Communications%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B12%26lt%3B%5C%2Fi%26gt%3B%281%29%2C%203127.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41467-021-23500-6%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41467-021-23500-6%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Cornelia%5Cu00a0de%20Lange%20syndrome-associated%20mutations%20cause%20a%20DNA%20damage%20signalling%20and%20repair%20defect%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Gabrielle%22%2C%22lastName%22%3A%22Olley%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Madapura%20M.%22%2C%22lastName%22%3A%22Pradeepa%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Graeme%20R.%22%2C%22lastName%22%3A%22Grimes%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Sandra%22%2C%22lastName%22%3A%22Piquet%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Sophie%20E.%22%2C%22lastName%22%3A%22Polo%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22David%20R.%22%2C%22lastName%22%3A%22FitzPatrick%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Wendy%20A.%22%2C%22lastName%22%3A%22Bickmore%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Charlene%22%2C%22lastName%22%3A%22Boumendil%22%7D%5D%2C%22abstractNote%22%3A%22Cornelia%20de%20Lange%20syndrome%20is%20a%20multisystem%20developmental%20disorder%20typically%20caused%20by%20mutations%20in%20the%20gene%20encoding%20the%20cohesin%20loader%20NIPBL.%20The%20associated%20phenotype%20is%20generally%20assumed%20to%20be%20the%20consequence%20of%20aberrant%20transcriptional%20regulation.%20Recently%2C%20we%20identified%20a%20missense%20mutation%20in%20BRD4%20associated%20with%20a%20Cornelia%20de%20Lange-like%20syndrome%20that%20reduces%20BRD4%20binding%20to%20acetylated%20histones.%20Here%20we%20show%20that%2C%20although%20this%20mutation%20reduces%20BRD4-occupancy%20at%20enhancers%20it%20does%20not%20affect%20transcription%20of%20the%20pluripotency%20network%20in%20mouse%20embryonic%20stem%20cells.%20Rather%2C%20it%20delays%20the%20cell%20cycle%2C%20increases%20DNA%20damage%20signalling%2C%20and%20perturbs%20regulation%20of%20DNA%20repair%20in%20mutant%20cells.%20This%20uncovers%20a%20role%20for%20BRD4%20in%20DNA%20repair%20pathway%20choice.%20Furthermore%2C%20we%20find%20evidence%20of%20a%20similar%20increase%20in%20DNA%20damage%20signalling%20in%20cells%20derived%20from%20NIPBL-deficient%20individuals%2C%20suggesting%20that%20defective%20DNA%20damage%20signalling%20and%20repair%20is%20also%20a%20feature%20of%20typical%20Cornelia%20de%20Lange%20syndrome.%22%2C%22date%22%3A%222021-05-25%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1038%5C%2Fs41467-021-23500-6%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%222041-1723%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A26%3A24Z%22%7D%7D%2C%7B%22key%22%3A%22N2CVDXA7%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Gonzalez-Estevez%20et%20al.%22%2C%22parsedDate%22%3A%222021-05-15%22%2C%22numChildren%22%3A3%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BGonzalez-Estevez%2C%20A.%2C%20Verrico%2C%20A.%2C%20Orniacki%2C%20C.%2C%20Reina-San-Martin%2C%20B.%2C%20%26amp%3B%20Doye%2C%20V.%20%282021%29.%20Integrity%20of%20the%20short%20arm%20of%20the%20nuclear%20pore%20Y-complex%20is%20required%20for%20mouse%20embryonic%20stem%20cell%20growth%20and%20differentiation.%20%26lt%3Bi%26gt%3BJournal%20of%20Cell%20Science%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B134%26lt%3B%5C%2Fi%26gt%3B%2810%29%2C%20jcs258340.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1242%5C%2Fjcs.258340%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1242%5C%2Fjcs.258340%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Integrity%20of%20the%20short%20arm%20of%20the%20nuclear%20pore%20Y-complex%20is%20required%20for%20mouse%20embryonic%20stem%20cell%20growth%20and%20differentiation%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alba%22%2C%22lastName%22%3A%22Gonzalez-Estevez%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Annalisa%22%2C%22lastName%22%3A%22Verrico%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Clarisse%22%2C%22lastName%22%3A%22Orniacki%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Bernardo%22%2C%22lastName%22%3A%22Reina-San-Martin%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22Many%20cellular%20processes%2C%20ranging%20from%20cell%20division%20to%20differentiation%2C%20are%20controlled%20by%20nuclear%20pore%20complexes%20%28NPCs%29.%20However%2C%20studying%20the%20contributions%20of%20individual%20NPC%20subunits%20to%20these%20processes%20in%20vertebrates%20has%20long%20been%20impeded%20by%20their%20complexity%20and%20the%20lack%20of%20efficient%20genetic%20tools.%20Here%2C%20we%20use%20genome%20editing%20in%20mouse%20embryonic%20stem%20cells%20%28mESCs%29%20to%20characterize%20the%20role%20of%20NPC%20structural%20components%2C%20focusing%20on%20the%20short%20arm%20of%20the%20Y-complex%20that%20comprises%20Nup85%2C%20Seh1%20and%20Nup43.%20We%20show%20that%20Seh1%20and%20Nup43%2C%20although%20dispensable%20in%20pluripotent%20mESCs%2C%20are%20required%20for%20their%20normal%20cell%20growth%20rates%2C%20their%20viability%20upon%20differentiation%20and%20for%20the%20maintenance%20of%20proper%20NPC%20density.%20mESCs%20with%20an%20N-terminally%20truncated%20Nup85%20mutation%20%28in%20which%20interaction%20with%20Seh1%20is%20greatly%20impaired%29%20feature%20a%20similar%20reduction%20of%20NPC%20density.%20However%2C%20their%20proliferation%20and%20differentiation%20are%20unaltered%2C%20indicating%20that%20it%20is%20the%20integrity%20of%20the%20Y-complex%2C%20rather%20than%20the%20number%20of%20NPCs%2C%20that%20is%20critical%20to%20ensure%20these%20processes.%22%2C%22date%22%3A%222021-05-15%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1242%5C%2Fjcs.258340%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221477-9137%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A26%3A24Z%22%7D%7D%2C%7B%22key%22%3A%228XJCA2PX%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Raich%20et%20al.%22%2C%22parsedDate%22%3A%222020-08-27%22%2C%22numChildren%22%3A2%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BRaich%2C%20N.%2C%20Contremoulins%2C%20V.%2C%20%26amp%3B%20Karess%2C%20R.%20E.%20%282020%29.%20Immunostaining%20of%20Whole-Mount%20Drosophila%20Testes%20for%203D%20Confocal%20Analysis%20of%20Large%20Spermatocytes.%20%26lt%3Bi%26gt%3BJournal%20of%20Visualized%20Experiments%3A%20JoVE%26lt%3B%5C%2Fi%26gt%3B%2C%20%28162%29.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.3791%5C%2F61061%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.3791%5C%2F61061%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Immunostaining%20of%20Whole-Mount%20Drosophila%20Testes%20for%203D%20Confocal%20Analysis%20of%20Large%20Spermatocytes%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Natacha%22%2C%22lastName%22%3A%22Raich%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Vincent%22%2C%22lastName%22%3A%22Contremoulins%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Roger%20E.%22%2C%22lastName%22%3A%22Karess%22%7D%5D%2C%22abstractNote%22%3A%22Drosophila%20testes%20are%20a%20powerful%20model%20system%20for%20studying%20biological%20processes%20including%20stem%20cell%20biology%2C%20nuclear%20architecture%2C%20meiosis%20and%20sperm%20development.%20However%2C%20immunolabeling%20of%20the%20whole%20Drosophila%20testis%20is%20often%20associated%20with%20significant%20non-uniformity%20of%20staining%20due%20to%20antibody%20penetration.%20Squashed%20preparations%20only%20partially%20overcome%20the%20problem%20since%20it%20decreases%20the%203D%20quality%20of%20the%20analyses.%20Herein%2C%20we%20describe%20a%20whole-mount%20protocol%20using%20NP40%20and%20heptane%20during%20fixation%20together%20with%20immunolabeling%20in%20liquid%20media.%20It%20preserves%20the%20volume%20suitable%20for%20confocal%20microscopy%20together%20with%20reproducible%20and%20reliable%20labeling.%20We%20show%20different%20examples%20of%203D%20reconstitution%20of%20spermatocyte%20nuclei%20from%20confocal%20sections.%20The%20intra-%20and%20inter-testes%20reproducibility%20allows%203D%20quantification%20and%20comparison%20of%20fluorescence%20between%20single%20cells%20from%20different%20genotypes.%20We%20used%20different%20components%20of%20the%20intranuclear%20MINT%20structure%20%28Mad1-containing%20Intra%20Nuclear%20Territory%29%20as%20well%20as%20two%20components%20associated%20with%20the%20nuclear%20pore%20complex%20to%20illustrate%20this%20protocol%20and%20its%20applications%20on%20the%20largest%20cells%20of%20the%20testis%2C%20the%20S4-S5%20spermatocytes.%22%2C%22date%22%3A%222020-08-27%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.3791%5C%2F61061%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221940-087X%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222023-07-31T14%3A57%3A02Z%22%7D%7D%2C%7B%22key%22%3A%22K737AHE4%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Cianciolo%20Cosentino%20et%20al.%22%2C%22parsedDate%22%3A%222020-02-19%22%2C%22numChildren%22%3A3%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BCianciolo%20Cosentino%2C%20C.%2C%20Berto%2C%20A.%2C%20Pelletier%2C%20S.%2C%20Hari%2C%20M.%2C%20Loffing%2C%20J.%2C%20Neuhauss%2C%20S.%20C.%20F.%2C%20%26amp%3B%20Doye%2C%20V.%20%282020%29.%20Publisher%20Correction%3A%20Moderate%20Nucleoporin%20133%20deficiency%20leads%20to%20glomerular%20damage%20in%20zebrafish.%20%26lt%3Bi%26gt%3BScientific%20Reports%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B10%26lt%3B%5C%2Fi%26gt%3B%281%29%2C%203326.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41598-020-58959-8%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41598-020-58959-8%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Publisher%20Correction%3A%20Moderate%20Nucleoporin%20133%20deficiency%20leads%20to%20glomerular%20damage%20in%20zebrafish%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Chiara%22%2C%22lastName%22%3A%22Cianciolo%20Cosentino%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alessandro%22%2C%22lastName%22%3A%22Berto%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22St%5Cu00e9phane%22%2C%22lastName%22%3A%22Pelletier%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Michelle%22%2C%22lastName%22%3A%22Hari%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Johannes%22%2C%22lastName%22%3A%22Loffing%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Stephan%20C.%20F.%22%2C%22lastName%22%3A%22Neuhauss%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22An%20amendment%20to%20this%20paper%20has%20been%20published%20and%20can%20be%20accessed%20via%20a%20link%20at%20the%20top%20of%20the%20paper.%22%2C%22date%22%3A%222020-02-19%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1038%5C%2Fs41598-020-58959-8%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%222045-2322%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A26%3A54Z%22%7D%7D%2C%7B%22key%22%3A%225GCFTI93%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Cianciolo%20Cosentino%20et%20al.%22%2C%22parsedDate%22%3A%222020-01-15%22%2C%22numChildren%22%3A3%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BCianciolo%20Cosentino%2C%20C.%2C%20Berto%2C%20A.%2C%20Pelletier%2C%20S.%2C%20Hari%2C%20M.%2C%20Loffing%2C%20J.%2C%20Neuhauss%2C%20S.%20C.%20F.%2C%20%26amp%3B%20Doye%2C%20V.%20%282020%29.%20Author%20Correction%3A%20Moderate%20Nucleoporin%20133%20deficiency%20leads%20to%20glomerular%20damage%20in%20zebrafish.%20%26lt%3Bi%26gt%3BScientific%20Reports%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B10%26lt%3B%5C%2Fi%26gt%3B%281%29%2C%20756.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41598-020-57829-7%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41598-020-57829-7%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Author%20Correction%3A%20Moderate%20Nucleoporin%20133%20deficiency%20leads%20to%20glomerular%20damage%20in%20zebrafish%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Chiara%22%2C%22lastName%22%3A%22Cianciolo%20Cosentino%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alessandro%22%2C%22lastName%22%3A%22Berto%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22St%5Cu00e9phane%22%2C%22lastName%22%3A%22Pelletier%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Michelle%22%2C%22lastName%22%3A%22Hari%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Johannes%22%2C%22lastName%22%3A%22Loffing%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Stephan%20C.%20F.%22%2C%22lastName%22%3A%22Neuhauss%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22An%20amendment%20to%20this%20paper%20has%20been%20published%20and%20can%20be%20accessed%20via%20a%20link%20at%20the%20top%20of%20the%20paper.%22%2C%22date%22%3A%222020-01-15%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1038%5C%2Fs41598-020-57829-7%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%222045-2322%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A27%3A07Z%22%7D%7D%2C%7B%22key%22%3A%2245SDNZ8J%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Menant%20and%20Karess%22%2C%22parsedDate%22%3A%222020%22%2C%22numChildren%22%3A2%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BMenant%2C%20A.%2C%20%26amp%3B%20Karess%2C%20R.%20E.%20%282020%29.%20Mutations%20in%20the%20Drosophila%20rough%20deal%20gene%20affecting%20RZZ%20kinetochore%20function.%20%26lt%3Bi%26gt%3BBiology%20of%20the%20Cell%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B112%26lt%3B%5C%2Fi%26gt%3B%2810%29%2C%20300%26%23×2013%3B315.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1111%5C%2Fboc.201900105%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1111%5C%2Fboc.201900105%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Mutations%20in%20the%20Drosophila%20rough%20deal%20gene%20affecting%20RZZ%20kinetochore%20function%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alexandra%22%2C%22lastName%22%3A%22Menant%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Roger%20E.%22%2C%22lastName%22%3A%22Karess%22%7D%5D%2C%22abstractNote%22%3A%22Background%20The%20RZZ%20complex%2C%20composed%20of%20the%20proteins%20Rough-Deal%20%28Rod%29%2C%20Zw10%20and%20Zwilch%2C%20plays%20a%20central%20role%20in%20the%20spindle%20assembly%20checkpoint%20%28SAC%29%2C%20which%20assures%20proper%20sister%20chromatid%20segregation%20during%20mitosis.%20RZZ%20contributes%20to%20the%20regulation%20of%20the%20spindle%20assembly%20checkpoint%20by%20helping%20to%20recruit%20Mad1%5Cu2013Mad2%20and%20the%20microtubule%20motor%20dynein%20to%20unattached%20kinetochores.%20It%20is%20an%20important%20component%20of%20the%20outer%20kinetochore%20and%20specifically%20the%20fibrous%20corona%20whose%20expansion%20is%20believed%20to%20facilitate%20microtubule%20capture.%20How%20RZZ%20carries%20out%20its%20diverse%20activities%20is%20only%20poorly%20understood.%20The%20C-terminal%20region%20of%20the%20Rod%20subunit%20is%20relatively%20well-conserved%20across%20metazoan%20phylogeny%2C%20but%20no%20function%20has%20been%20attributed%20to%20it.%20Results%20To%20explore%20the%20importance%20of%20the%20Rod_C%20domain%20in%20RZZ%20function%20in%20Drosophila%2C%20we%20generated%20a%20series%20of%20point%20mutations%20in%20a%20stretch%20of%20200%20residues%20within%20this%20domain%20and%20we%20report%20here%20their%20phenotypes.%20Several%20of%20the%20mutations%20profoundly%20disrupt%20recruitment%20of%20RZZ%20to%20kinetochores%2C%20including%20one%20in%20a%20temperature-sensitive%20manner%2C%20while%20still%20retaining%20the%20capacity%20to%20assemble%20into%20a%20complex%20with%20Zw10%20and%20Zwilch.%20Others%20affect%20aspects%20of%20dynein%20activity%20or%20recruitment%20at%20the%20kinetochore.%20Conclusions%20and%20Significance%20These%20results%20suggest%20that%20the%20Rod_C%20domain%20participates%20in%20the%20protein%20interactions%20necessary%20for%20RZZ%20recruitment%20and%20functionality%20at%20kinetochores.%22%2C%22date%22%3A%222020%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1111%5C%2Fboc.201900105%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22https%3A%5C%2F%5C%2Fonlinelibrary.wiley.com%5C%2Fdoi%5C%2Fabs%5C%2F10.1111%5C%2Fboc.201900105%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221768-322X%22%2C%22language%22%3A%22en%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222023-05-23T12%3A10%3A59Z%22%7D%7D%2C%7B%22key%22%3A%22GN6B5RFE%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Cianciolo%20Cosentino%20et%20al.%22%2C%22parsedDate%22%3A%222019-03-18%22%2C%22numChildren%22%3A3%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BCianciolo%20Cosentino%2C%20C.%2C%20Berto%2C%20A.%2C%20Pelletier%2C%20S.%2C%20Hari%2C%20M.%2C%20Loffing%2C%20J.%2C%20Neuhauss%2C%20S.%20C.%20F.%2C%20%26amp%3B%20Doye%2C%20V.%20%282019%29.%20Moderate%20Nucleoporin%20133%20deficiency%20leads%20to%20glomerular%20damage%20in%20zebrafish.%20%26lt%3Bi%26gt%3BScientific%20Reports%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B9%26lt%3B%5C%2Fi%26gt%3B%281%29%2C%204750.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41598-019-41202-4%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1038%5C%2Fs41598-019-41202-4%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Moderate%20Nucleoporin%20133%20deficiency%20leads%20to%20glomerular%20damage%20in%20zebrafish%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Chiara%22%2C%22lastName%22%3A%22Cianciolo%20Cosentino%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alessandro%22%2C%22lastName%22%3A%22Berto%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22St%5Cu00e9phane%22%2C%22lastName%22%3A%22Pelletier%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Michelle%22%2C%22lastName%22%3A%22Hari%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Johannes%22%2C%22lastName%22%3A%22Loffing%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Stephan%20C.%20F.%22%2C%22lastName%22%3A%22Neuhauss%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22Although%20structural%20nuclear%20pore%20proteins%20%28nucleoporins%29%20are%20seemingly%20required%20in%20every%20cell%20type%20to%20assemble%20a%20functional%20nuclear%20transport%20machinery%2C%20mutations%20or%20deregulation%20of%20a%20subset%20of%20them%20have%20been%20associated%20with%20specific%20human%20hereditary%20diseases.%20In%20particular%2C%20previous%20genetic%20studies%20of%20patients%20with%20nephrotic%20syndrome%20identified%20mutations%20in%20Nup107%20that%20impaired%20the%20expression%20or%20the%20localization%20of%20its%20direct%20partner%20at%20nuclear%20pores%2C%20Nup133.%20In%20the%20present%20study%2C%20we%20characterized%20the%20zebrafish%20nup133%20orthologous%20gene%20and%20its%20expression%20pattern%20during%20larval%20development.%20Using%20a%20morpholino-mediated%20gene%20knockdown%2C%20we%20show%20that%20partial%20depletion%20of%20Nup133%20in%20zebrafish%20larvae%20leads%20to%20the%20formation%20of%20kidney%20cysts%2C%20a%20phenotype%20that%20can%20be%20rescued%20by%20co-injection%20of%20wild%20type%20mRNA.%20Analysis%20of%20different%20markers%20for%20tubular%20and%20glomerular%20development%20shows%20that%20the%20overall%20kidney%20development%20is%20not%20affected%20by%20nup133%20knockdown.%20Likewise%2C%20no%20gross%20defect%20in%20nuclear%20pore%20complex%20assembly%20was%20observed%20in%20these%20nup133%20morphants.%20On%20the%20other%20hand%2C%20nup133%20downregulation%20results%20in%20proteinuria%20and%20moderate%20foot%20process%20effacement%2C%20mimicking%20some%20of%20the%20abnormalities%20typically%20featured%20by%20patients%20with%20nephrotic%20syndrome.%20These%20data%20indicate%20that%20nup133%20is%20a%20new%20gene%20required%20for%20proper%20glomerular%20structure%20and%20function%20in%20zebrafish.%22%2C%22date%22%3A%222019-03-18%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1038%5C%2Fs41598-019-41202-4%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%222045-2322%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A27%3A23Z%22%7D%7D%2C%7B%22key%22%3A%22HHSPUI9D%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22creatorSummary%22%3A%22Raich%20et%20al.%22%2C%22parsedDate%22%3A%222018-10-17%22%2C%22numChildren%22%3A1%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BRaich%2C%20N.%2C%20Mahmoudi%2C%20S.%2C%20Emre%2C%20D.%2C%20%26amp%3B%20Karess%2C%20R.%20E.%20%282018%29.%20Mad1%20influences%20interphase%20nucleoplasm%20organization%20and%20chromatin%20regulation%20in%20Drosophila.%20%26lt%3Bi%26gt%3BOpen%20Biology%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B8%26lt%3B%5C%2Fi%26gt%3B%2810%29%2C%20180166.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1098%5C%2Frsob.180166%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1098%5C%2Frsob.180166%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Mad1%20influences%20interphase%20nucleoplasm%20organization%20and%20chromatin%20regulation%20in%20Drosophila%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Natacha%22%2C%22lastName%22%3A%22Raich%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Souhir%22%2C%22lastName%22%3A%22Mahmoudi%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Doruk%22%2C%22lastName%22%3A%22Emre%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Roger%20E.%22%2C%22lastName%22%3A%22Karess%22%7D%5D%2C%22abstractNote%22%3A%22The%20Drosophila%20Mad1%20spindle%20checkpoint%20protein%20helps%20organize%20several%20nucleoplasmic%20components%2C%20and%20flies%20lacking%20Mad1%20present%20changes%20in%20gene%20expression%20reflecting%20altered%20chromatin%20conformation.%20In%20interphase%2C%20checkpoint%20protein%20Mad1%20is%20usually%20described%20as%20localizing%20to%20the%20inner%20nuclear%20envelope%20by%20binding%20the%20nucleoporin%20Tpr%2C%20an%20interaction%20believed%20to%20contribute%20to%20proper%20mitotic%20regulation.%20Whether%20Mad1%20has%20other%20nuclear%20interphase%20functions%20is%20unknown.%20We%20found%20in%20Drosophila%20that%20Mad1%20is%20present%20in%20nuclei%20of%20both%20mitotic%20and%20postmitotic%20tissues.%20Three%20proteins%20implicated%20in%20various%20aspects%20of%20chromatin%20organization%20co-immunoprecipitated%20with%20Mad1%20from%20fly%20embryos%3A%20Mtor%5C%2FTpr%2C%20the%20SUMO%20peptidase%20Ulp1%20and%20Raf2%2C%20a%20subunit%20of%20a%20Polycomb-like%20complex.%20In%20primary%20spermatocytes%2C%20all%20four%20proteins%20colocalized%20in%20a%20previously%20undescribed%20chromatin-associated%20structure%20called%20here%20a%20MINT%20%28Mad1-containing%20IntraNuclear%20Territory%29.%20MINT%20integrity%20required%20all%20four%20proteins.%20In%20mad1%20mutant%20spermatocytes%2C%20the%20other%20proteins%20were%20no%20longer%20confined%20to%20chromatin%20domains%20but%20instead%20dispersed%20throughout%20the%20nucleoplasm.%20mad1%20flies%20also%20presented%20phenotypes%20indicative%20of%20excessive%20chromatin%20of%20heterochromatic%20character%20during%20development%20of%20somatic%20tissues.%20Together%20these%20results%20suggest%20that%20Drosophila%20Mad1%2C%20by%20helping%20organize%20its%20interphase%20protein%20partners%20in%20the%20nucleoplasm%2C%20contributes%20to%20proper%20chromatin%20regulation.%22%2C%22date%22%3A%222018-10-17%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1098%5C%2Frsob.180166%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22https%3A%5C%2F%5C%2Froyalsocietypublishing.org%5C%2Fdoi%5C%2F10.1098%5C%2Frsob.180166%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%22%22%2C%22language%22%3A%22%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222023-07-31T14%3A57%3A32Z%22%7D%7D%2C%7B%22key%22%3A%22PE9BHLD9%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Souquet%20et%20al.%22%2C%22parsedDate%22%3A%222018-05-22%22%2C%22numChildren%22%3A3%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BSouquet%2C%20B.%2C%20Freed%2C%20E.%2C%20Berto%2C%20A.%2C%20Andric%2C%20V.%2C%20Audug%26%23xE9%3B%2C%20N.%2C%20Reina-San-Martin%2C%20B.%2C%20Lacy%2C%20E.%2C%20%26amp%3B%20Doye%2C%20V.%20%282018%29.%20Nup133%20Is%20Required%20for%20Proper%20Nuclear%20Pore%20Basket%20Assembly%20and%20Dynamics%20in%20Embryonic%20Stem%20Cells.%20%26lt%3Bi%26gt%3BCell%20Reports%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B23%26lt%3B%5C%2Fi%26gt%3B%288%29%2C%202443%26%23×2013%3B2454.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.celrep.2018.04.070%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.celrep.2018.04.070%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Nup133%20Is%20Required%20for%20Proper%20Nuclear%20Pore%20Basket%20Assembly%20and%20Dynamics%20in%20Embryonic%20Stem%20Cells%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Benoit%22%2C%22lastName%22%3A%22Souquet%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Ellen%22%2C%22lastName%22%3A%22Freed%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alessandro%22%2C%22lastName%22%3A%22Berto%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Vedrana%22%2C%22lastName%22%3A%22Andric%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Nicolas%22%2C%22lastName%22%3A%22Audug%5Cu00e9%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Bernardo%22%2C%22lastName%22%3A%22Reina-San-Martin%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Elizabeth%22%2C%22lastName%22%3A%22Lacy%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22Nup133%20belongs%20to%20the%20Y-complex%2C%20a%20key%20component%20of%20the%20nuclear%20pore%20complex%20%28NPC%29%20scaffold.%20Studies%20on%20a%20null%20mutation%20in%20mice%20previously%20revealed%20that%20Nup133%20is%20essential%20for%20embryonic%20development%20but%5Cu00a0not%20for%20mouse%20embryonic%20stem%20cell%20%28mESC%29%20proliferation.%20Using%20single-pore%20detection%20and%20average%20NE-fluorescence%20intensity%2C%20we%20find%20that%20Nup133%20is%20dispensable%20for%20interphase%20and%20postmitotic%20NPC%20scaffold%20assembly%20in%20pluripotent%20mESCs.%20However%2C%20loss%20of%20Nup133%20specifically%20perturbs%20the%20formation%20of%20the%20nuclear%20basket%20as%20manifested%20by%20the%20absence%20of%20Tpr%20in%20about%20half%20of%20the%20NPCs%20combined%20with%20altered%20dynamics%20of%20Nup153.%20We%20further%20demonstrate%20that%20its%20central%20domain%20mediates%20Nup133%26%23039%3Bs%20role%20in%20assembling%20Tpr%20and%20Nup153%20into%20a%20properly%20configured%20nuclear%20basket.%20Our%20findings%20thus%20revisit%20the%20role%20of%20the%20Y-complex%20in%20pore%20biogenesis%20and%20provide%20insights%20into%20the%20interplay%20between%20NPC%20scaffold%20architecture%2C%20nuclear%20basket%20assembly%2C%20and%20the%20generation%20of%20heterogeneity%20among%20NPCs.%22%2C%22date%22%3A%222018-05-22%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1016%5C%2Fj.celrep.2018.04.070%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%222211-1247%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A27%3A49Z%22%7D%7D%2C%7B%22key%22%3A%22E7759ARZ%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Berto%20et%20al.%22%2C%22parsedDate%22%3A%222018-05%22%2C%22numChildren%22%3A4%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BBerto%2C%20A.%2C%20Yu%2C%20J.%2C%20Morchoisne-Bolhy%2C%20S.%2C%20Bertipaglia%2C%20C.%2C%20Vallee%2C%20R.%2C%20Dumont%2C%20J.%2C%20Ochsenbein%2C%20F.%2C%20Guerois%2C%20R.%2C%20%26amp%3B%20Doye%2C%20V.%20%282018%29.%20Disentangling%20the%20molecular%20determinants%20for%20Cenp-F%20localization%20to%20nuclear%20pores%20and%20kinetochores.%20%26lt%3Bi%26gt%3BEMBO%20Reports%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B19%26lt%3B%5C%2Fi%26gt%3B%285%29%2C%20e44742.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.15252%5C%2Fembr.201744742%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.15252%5C%2Fembr.201744742%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Disentangling%20the%20molecular%20determinants%20for%20Cenp-F%20localization%20to%20nuclear%20pores%20and%20kinetochores%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alessandro%22%2C%22lastName%22%3A%22Berto%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Jinchao%22%2C%22lastName%22%3A%22Yu%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22St%5Cu00e9phanie%22%2C%22lastName%22%3A%22Morchoisne-Bolhy%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Chiara%22%2C%22lastName%22%3A%22Bertipaglia%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Richard%22%2C%22lastName%22%3A%22Vallee%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Julien%22%2C%22lastName%22%3A%22Dumont%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Francoise%22%2C%22lastName%22%3A%22Ochsenbein%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Raphael%22%2C%22lastName%22%3A%22Guerois%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22Cenp-F%20is%20a%20multifaceted%20protein%20implicated%20in%20cancer%20and%20developmental%20pathologies.%20The%20Cenp-F%20C-terminal%20region%20contains%20overlapping%20binding%20sites%20for%20numerous%20proteins%20that%20contribute%20to%20its%20functions%20throughout%20the%20cell%20cycle.%20Here%2C%20we%20focus%20on%20the%20nuclear%20pore%20protein%20Nup133%20that%20interacts%20with%20Cenp-F%20both%20at%20nuclear%20pores%20in%20prophase%20and%20at%20kinetochores%20in%20mitosis%2C%20and%20on%20the%20kinase%20Bub1%2C%20known%20to%20contribute%20to%20Cenp-F%20targeting%20to%20kinetochores.%20By%20combining%20in%20silico%20structural%20modeling%20and%20yeast%20two-hybrid%20assays%2C%20we%20generate%20an%20interaction%20model%20between%20a%20conserved%20helix%20within%20the%20Nup133%20%5Cu03b2-propeller%20and%20a%20short%20leucine%20zipper-containing%20dimeric%20segment%20of%20Cenp-F.%20We%20thereby%20create%20mutants%20affecting%20the%20Nup133%5C%2FCenp-F%20interface%20and%20show%20that%20they%20prevent%20Cenp-F%20localization%20to%20the%20nuclear%20envelope%2C%20but%20not%20to%20kinetochores.%20Conversely%2C%20a%20point%20mutation%20within%20an%20adjacent%20leucine%20zipper%20affecting%20the%20kinetochore%20targeting%20of%20Cenp-F%20KT-core%20domain%20impairs%20its%20interaction%20with%20Bub1%2C%20but%20not%20with%20Nup133%2C%20identifying%20Bub1%20as%20the%20direct%20KT-core%20binding%20partner%20of%20Cenp-F.%20Finally%2C%20we%20show%20that%20Cenp-E%20redundantly%20contributes%20together%20with%20Bub1%20to%20the%20recruitment%20of%20Cenp-F%20to%20kinetochores.%22%2C%22date%22%3A%222018-05%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.15252%5C%2Fembr.201744742%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221469-3178%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A27%3A49Z%22%7D%7D%2C%7B%22key%22%3A%22L6RJID2H%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Berto%20and%20Doye%22%2C%22parsedDate%22%3A%222018%22%2C%22numChildren%22%3A3%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BBerto%2C%20A.%2C%20%26amp%3B%20Doye%2C%20V.%20%282018%29.%20Regulation%20of%20Cenp-F%20localization%20to%20nuclear%20pores%20and%20kinetochores.%20%26lt%3Bi%26gt%3BCell%20Cycle%20%28Georgetown%2C%20Tex.%29%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B17%26lt%3B%5C%2Fi%26gt%3B%2817%29%2C%202122%26%23×2013%3B2133.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1080%5C%2F15384101.2018.1520569%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1080%5C%2F15384101.2018.1520569%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Regulation%20of%20Cenp-F%20localization%20to%20nuclear%20pores%20and%20kinetochores%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Alessandro%22%2C%22lastName%22%3A%22Berto%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%5D%2C%22abstractNote%22%3A%22In%20metazoans%2C%20the%20assembly%20of%20kinetochores%20on%20centrometric%20chromatin%20and%20the%20dismantling%20of%20nuclear%20pore%20complexes%20are%20processes%20that%20have%20to%20be%20tightly%20coordinated%20to%20ensure%20the%20proper%20assembly%20of%20the%20mitotic%20spindle%20and%20a%20successful%20mitosis.%20It%20is%20therefore%20noteworthy%20that%20these%20two%20macromolecular%20assemblies%20share%20a%20subset%20of%20constituents.%20One%20of%20these%20multifaceted%20components%20is%20Cenp-F%2C%20a%20protein%20implicated%20in%20cancer%20and%20developmental%20pathologies.%20During%20the%20cell%20cycle%2C%20Cenp-F%20localizes%20in%20multiple%20cellular%20structures%20including%20the%20nuclear%20envelope%20in%20late%20G2%5C%2Fearly%20prophase%20and%20kinetochores%20throughout%20mitosis.%20We%20recently%20characterized%20the%20molecular%20determinants%20of%20Cenp-F%20interaction%20with%20Nup133%2C%20a%20structural%20nuclear%20pore%20constituent.%20In%20parallel%20with%20two%20other%20independent%20studies%2C%20we%20further%20elucidated%20the%20mechanisms%20governing%20Cenp-F%20kinetochore%20recruitment%20that%20mainly%20relies%20on%20its%20interaction%20with%20Bub1%2C%20with%20redundant%20contribution%20of%20Cenp-E%20upon%20acute%20microtubule%20depolymerisation.%20Here%20we%20synthesize%20the%20current%20literature%20regarding%20the%20dual%20location%20of%20Cenp-F%20at%20nuclear%20pores%20and%20kinetochores%20and%20extend%20our%20discussion%20to%20the%20regulation%20of%20these%20NPC%20and%20kinetochore%20localizations%20by%20mitotic%20kinase%20and%20spindle%20microtubules.%22%2C%22date%22%3A%222018%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1080%5C%2F15384101.2018.1520569%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221551-4005%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A27%3A35Z%22%7D%7D%2C%7B%22key%22%3A%22L3Z9VEXR%22%2C%22library%22%3A%7B%22id%22%3A2913254%7D%2C%22meta%22%3A%7B%22lastModifiedByUser%22%3A%7B%22id%22%3A11274337%2C%22username%22%3A%22Charlotte_Brancaz%22%2C%22name%22%3A%22%22%2C%22links%22%3A%7B%22alternate%22%3A%7B%22href%22%3A%22https%3A%5C%2F%5C%2Fwww.zotero.org%5C%2Fcharlotte_brancaz%22%2C%22type%22%3A%22text%5C%2Fhtml%22%7D%7D%7D%2C%22creatorSummary%22%3A%22Martino%20et%20al.%22%2C%22parsedDate%22%3A%222017-10-23%22%2C%22numChildren%22%3A5%7D%2C%22bib%22%3A%22%26lt%3Bdiv%20class%3D%26quot%3Bcsl-bib-body%26quot%3B%20style%3D%26quot%3Bline-height%3A%202%3B%20padding-left%3A%201em%3B%20text-indent%3A-1em%3B%26quot%3B%26gt%3B%5Cn%20%20%26lt%3Bdiv%20class%3D%26quot%3Bcsl-entry%26quot%3B%26gt%3BMartino%2C%20L.%2C%20Morchoisne-Bolhy%2C%20S.%2C%20Cheerambathur%2C%20D.%20K.%2C%20Van%20Hove%2C%20L.%2C%20Dumont%2C%20J.%2C%20Joly%2C%20N.%2C%20Desai%2C%20A.%2C%20Doye%2C%20V.%2C%20%26amp%3B%20Pintard%2C%20L.%20%282017%29.%20Channel%20Nucleoporins%20Recruit%20PLK-1%20to%20Nuclear%20Pore%20Complexes%20to%20Direct%20Nuclear%20Envelope%20Breakdown%20in%20C.%26%23xA0%3Belegans.%20%26lt%3Bi%26gt%3BDevelopmental%20Cell%26lt%3B%5C%2Fi%26gt%3B%2C%20%26lt%3Bi%26gt%3B43%26lt%3B%5C%2Fi%26gt%3B%282%29%2C%20157-171.e7.%20%26lt%3Ba%20class%3D%26%23039%3Bzp-DOIURL%26%23039%3B%20href%3D%26%23039%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.devcel.2017.09.019%26%23039%3B%26gt%3Bhttps%3A%5C%2F%5C%2Fdoi.org%5C%2F10.1016%5C%2Fj.devcel.2017.09.019%26lt%3B%5C%2Fa%26gt%3B%26lt%3B%5C%2Fdiv%26gt%3B%5Cn%26lt%3B%5C%2Fdiv%26gt%3B%22%2C%22data%22%3A%7B%22itemType%22%3A%22journalArticle%22%2C%22title%22%3A%22Channel%20Nucleoporins%20Recruit%20PLK-1%20to%20Nuclear%20Pore%20Complexes%20to%20Direct%20Nuclear%20Envelope%20Breakdown%20in%20C.%5Cu00a0elegans%22%2C%22creators%22%3A%5B%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Lisa%22%2C%22lastName%22%3A%22Martino%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22St%5Cu00e9phanie%22%2C%22lastName%22%3A%22Morchoisne-Bolhy%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Dhanya%20K.%22%2C%22lastName%22%3A%22Cheerambathur%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Lucie%22%2C%22lastName%22%3A%22Van%20Hove%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Julien%22%2C%22lastName%22%3A%22Dumont%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Nicolas%22%2C%22lastName%22%3A%22Joly%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Arshad%22%2C%22lastName%22%3A%22Desai%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Val%5Cu00e9rie%22%2C%22lastName%22%3A%22Doye%22%7D%2C%7B%22creatorType%22%3A%22author%22%2C%22firstName%22%3A%22Lionel%22%2C%22lastName%22%3A%22Pintard%22%7D%5D%2C%22abstractNote%22%3A%22In%20animal%20cells%2C%20nuclear%20envelope%20breakdown%20%28NEBD%29%20is%20required%20for%20proper%20chromosome%20segregation.%20Whereas%20mitotic%20kinases%20have%20been%20implicated%20in%20NEBD%2C%20how%20they%20coordinate%20their%20activity%20to%20trigger%20this%20event%20is%20unclear.%20Here%2C%20we%20show%20that%20both%20in%20human%20cells%20and%20Caenorhabditis%20elegans%2C%20the%20Polo-like%20kinase%201%20%28PLK-1%29%20is%20recruited%20to%20the%20nuclear%20pore%20complexes%2C%20just%20prior%20to%20NEBD%2C%20through%20its%20Polo-box%20domain%20%28PBD%29.%20We%20provide%20evidence%20that%20PLK-1%20localization%20to%20the%20nuclear%20envelope%20%28NE%29%20is%20required%20for%20efficient%20NEBD.%20We%20identify%20the%20central%20channel%5Cu00a0nucleoporins%20NPP-1%5C%2FNup58%2C%20NPP-4%5C%2FNup54%2C%20and%20NPP-11%5C%2FNup62%20as%20the%20critical%20factors%20anchoring%20PLK-1%20to%20the%20NE%20in%20C.%5Cu00a0elegans.%20In%20particular%2C%20NPP-1%2C%20NPP-4%2C%20and%20NPP-11%20primed%20at%20multiple%20Polo-docking%20sites%20by%20Cdk1%20and%20PLK-1%20itself%20physically%20interact%20with%20the%20PLK-1%20PBD.%20We%20conclude%20that%20nucleoporins%20play%20an%20unanticipated%20regulatory%20role%20in%20NEBD%2C%20by%20recruiting%20PLK-1%20to%20the%20NE%20thereby%20facilitating%20phosphorylation%20of%20critical%20downstream%20targets.%22%2C%22date%22%3A%222017-10-23%22%2C%22section%22%3A%22%22%2C%22partNumber%22%3A%22%22%2C%22partTitle%22%3A%22%22%2C%22DOI%22%3A%2210.1016%5C%2Fj.devcel.2017.09.019%22%2C%22citationKey%22%3A%22%22%2C%22url%22%3A%22%22%2C%22PMID%22%3A%22%22%2C%22PMCID%22%3A%22%22%2C%22ISSN%22%3A%221878-1551%22%2C%22language%22%3A%22eng%22%2C%22collections%22%3A%5B%22YZUD6P7F%22%5D%2C%22dateModified%22%3A%222022-09-05T13%3A28%3A03Z%22%7D%7D%5D%7D
Vadrot, N., Moulin, M., Ferreiro, A., Richard, P., & Buendia, B. (2026). LAP2 Isoform Profile in Heart Ageing and in Cardiac Cell Proliferation and Differentiation: Input From CRISPR-Cas9-mediated LAP2a Knockdown in H9C2.
International Journal of Medical Sciences,
23(3), 741–757.
https://doi.org/10.7150/ijms.114095
Taniguchi, R., Orniacki, C., Kreysing, J. P., Zila, V., Zimmerli, C. E., Böhm, S., Turoňová, B., Kräusslich, H.-G., Doye, V., & Beck, M. (2025). Nuclear pores safeguard the integrity of the nuclear envelope.
Nature Cell Biology, 1–14.
https://doi.org/10.1038/s41556-025-01648-3
Dultz, E., & Doye, V. (2025). Opening the gate: Complexity and modularity of the nuclear pore scaffold and basket.
Current Opinion in Cell Biology,
92, 102461.
https://doi.org/10.1016/j.ceb.2024.102461
Nobari, P., Doye, V., & Boumendil, C. (2023). Metazoan nuclear pore complexes in gene regulation and genome stability.
DNA Repair,
130, 103565.
https://doi.org/10.1016/j.dnarep.2023.103565
Orniacki, C., Verrico, A., Pelletier, S., Souquet, B., Coulpier, F., Jourdren, L., Benetti, S., & Doye, V. (2023). Y-complex nucleoporins independently contribute to nuclear pore assembly and gene regulation in neuronal progenitors.
Journal of Cell Science, jcs.261151.
https://doi.org/10.1242/jcs.261151
Bouglita, W., Rabhi, S., Raich, N., Bouabid, C., Belghith, C., Slimani, O., Hkimi, C., Ghedira, K., Karess, R. E., Guizani-Tabbane, L., Attia, L., & Rabhi, I. (2022). Microbiological and molecular screening of Candida spp. isolated from genital tract of asymptomatic pregnant women.
Journal of Medical Microbiology,
71(9), 001589.
https://doi.org/10.1099/jmm.0.001589
Olley, G., Pradeepa, M. M., Grimes, G. R., Piquet, S., Polo, S. E., FitzPatrick, D. R., Bickmore, W. A., & Boumendil, C. (2021). Cornelia de Lange syndrome-associated mutations cause a DNA damage signalling and repair defect.
Nature Communications,
12(1), 3127.
https://doi.org/10.1038/s41467-021-23500-6
Gonzalez-Estevez, A., Verrico, A., Orniacki, C., Reina-San-Martin, B., & Doye, V. (2021). Integrity of the short arm of the nuclear pore Y-complex is required for mouse embryonic stem cell growth and differentiation.
Journal of Cell Science,
134(10), jcs258340.
https://doi.org/10.1242/jcs.258340
Raich, N., Contremoulins, V., & Karess, R. E. (2020). Immunostaining of Whole-Mount Drosophila Testes for 3D Confocal Analysis of Large Spermatocytes.
Journal of Visualized Experiments: JoVE, (162).
https://doi.org/10.3791/61061
Cianciolo Cosentino, C., Berto, A., Pelletier, S., Hari, M., Loffing, J., Neuhauss, S. C. F., & Doye, V. (2020). Publisher Correction: Moderate Nucleoporin 133 deficiency leads to glomerular damage in zebrafish.
Scientific Reports,
10(1), 3326.
https://doi.org/10.1038/s41598-020-58959-8
Cianciolo Cosentino, C., Berto, A., Pelletier, S., Hari, M., Loffing, J., Neuhauss, S. C. F., & Doye, V. (2020). Author Correction: Moderate Nucleoporin 133 deficiency leads to glomerular damage in zebrafish.
Scientific Reports,
10(1), 756.
https://doi.org/10.1038/s41598-020-57829-7
Menant, A., & Karess, R. E. (2020). Mutations in the Drosophila rough deal gene affecting RZZ kinetochore function.
Biology of the Cell,
112(10), 300–315.
https://doi.org/10.1111/boc.201900105
Cianciolo Cosentino, C., Berto, A., Pelletier, S., Hari, M., Loffing, J., Neuhauss, S. C. F., & Doye, V. (2019). Moderate Nucleoporin 133 deficiency leads to glomerular damage in zebrafish.
Scientific Reports,
9(1), 4750.
https://doi.org/10.1038/s41598-019-41202-4
Raich, N., Mahmoudi, S., Emre, D., & Karess, R. E. (2018). Mad1 influences interphase nucleoplasm organization and chromatin regulation in Drosophila.
Open Biology,
8(10), 180166.
https://doi.org/10.1098/rsob.180166
Souquet, B., Freed, E., Berto, A., Andric, V., Audugé, N., Reina-San-Martin, B., Lacy, E., & Doye, V. (2018). Nup133 Is Required for Proper Nuclear Pore Basket Assembly and Dynamics in Embryonic Stem Cells.
Cell Reports,
23(8), 2443–2454.
https://doi.org/10.1016/j.celrep.2018.04.070
Berto, A., Yu, J., Morchoisne-Bolhy, S., Bertipaglia, C., Vallee, R., Dumont, J., Ochsenbein, F., Guerois, R., & Doye, V. (2018). Disentangling the molecular determinants for Cenp-F localization to nuclear pores and kinetochores.
EMBO Reports,
19(5), e44742.
https://doi.org/10.15252/embr.201744742
Berto, A., & Doye, V. (2018). Regulation of Cenp-F localization to nuclear pores and kinetochores.
Cell Cycle (Georgetown, Tex.),
17(17), 2122–2133.
https://doi.org/10.1080/15384101.2018.1520569
Martino, L., Morchoisne-Bolhy, S., Cheerambathur, D. K., Van Hove, L., Dumont, J., Joly, N., Desai, A., Doye, V., & Pintard, L. (2017). Channel Nucleoporins Recruit PLK-1 to Nuclear Pore Complexes to Direct Nuclear Envelope Breakdown in C. elegans.
Developmental Cell,
43(2), 157-171.e7.
https://doi.org/10.1016/j.devcel.2017.09.019
Theses
Clarisse Orniacki (01/10/2018 – 12/12/2022) – Université Paris Cité.
[ les nucléoporines du complexe Y dans la différenciation des cellules souches embryonnaires: impact sur l’intégrité des pores nucléaires et la régulation des gènes]
Y-complex nucleoporins in embryonic stem cell differentiation; impact on nuclear pore integrity and gene regulation
Alba Gonzalez-Estevez (01/10/2016 – 19/12/2019) – Université Paris Diderot
[À la frontière entre division et différenciation cellulaire : rôle de Seh1 dans les cellules souches embryonnaires de souris]
On the edge between cell division and cell differentiation: role of Seh1 in mouse embryonic stem cells
Alessandro Berto (01/10/2013 – 11/12/2017) – Université Paris-Saclay
[Implication de la sous-unité Nup133 des pores nucléaires dans la division et la différenciation cellulaires : partenaires et mécanismes]
Implication of the Nup133 subunit of nuclear pores in cell division and differentiation: partners and mechanisms
Aurélie Floch (01/10/2010 – 26/09/2014) – Université Paris-Sud
Mécanismes d’adressage de Pom33, protéine transmembranaire associée aux pores nucléaires chez la levure Saccharomyces cerevisiae
Anne CHADRIN (01/10/2006 – 02/07/2010) -Université Paris XI
Analyses fonctionnelles d’une nouvelle protéine de l’enveloppe nucléaire chez la levure Saccharomyces cerevisiae
Michela ZUCCOLO (01/09/2005- 19/07/2008) -Université Paris XI
Implication du complexe Nup107-160 des pores nucléaires dans l’entrée, la progression et la sortie de mitose.
Annabelle Alves (01/09/2001- 01/07/2005) -Université Paris XI- Orsay
Analyse fonctionnelle du complexe NUP107-160 des pores nucléaires au cours du cycle cellulaire des vertébrés
Siau Wei BAï (01/09/2000 – 10/12/2004)- Université Paris XI- Orsay
Caractérisation fonctionnele du complexe Nup107: un sous-complexe protéique du pore nucléaire chez la levure Schizoszaccharomyces pombe
Naïma BELGAREH (01/09/ 1995- 27/09/1999) – Université Paris VI
Etude de l’assemblage et de la dynamique des pores nucléaires
Collaborations
Ongoing collaborations:
- Martin BECK, Max Planck Insitute of Biophysics, Frankfurt, Germany
- Corinne ANTIGNAC et Géraldine MOLLET, Institut Image, Paris, France;
- Brian BURKE, IMB, A*Star, Singapore;
- Andreas SCHEDL, Institut de Biologie Valrose, Nice, France;
- Fanny COULPIER and Laurent JOURDREN, Genomics facility, IBENS, Paris, France
Former collaborations:
- Johannes LOFFING and Stephan C. F. NEUHAUSS, University of Zurich, Switzerland;
- Bernardo REINA−SAN−MARTIN, IGBMC, Illkirch, France;
- Elizabeth LACY, Memorial Sloan Kettering Cancer Center, New York, USA;
- Raphael GUEROIS, Institut de Biologie et de Technologies de Saclay (iBiTec-S), Gif-sur-Yvette, France;
- Richard VALLEE – Department of Pathology and Cell Biology, Columbia University, New York, USA;
- Julien DUMONT, Institut Jacques-Monod IJM, Paris, France;
- Lionel PINTARD, Institut Jacques-Monod IJM, Paris, France
Grants - Funding
The team has been recognized as “équipe FRM 2020 ” by the Fondation pour la Recherche Médicale and was supported in 2019 by a grant form IDEX Université de Paris. The team has been supported by the Labex “Who am I?” and ANR, and currently hosts in its lab-space the members of the enSCORE facility, founded by the Labex “Who am I?”and dedicated to the development of neuronal organoids.



News
Job Offer



 Brigitte BUENDIA, Researcher, DOYE LAB+33 (0)1 57 27 80 60, room 344B
Brigitte BUENDIA, Researcher, DOYE LAB+33 (0)1 57 27 80 60, room 344B Marie DIDELOT, Intern, DOYE LAB+33 (0)1 57 27 80 62, room 314B
Marie DIDELOT, Intern, DOYE LAB+33 (0)1 57 27 80 62, room 314B Roger KARESS, Emeritus researcher, DOYE LAB+33 (0)1 57 27 80 95, room 344B
Roger KARESS, Emeritus researcher, DOYE LAB+33 (0)1 57 27 80 95, room 344B Fernanda LOPEZ GARCIA, Intern, DOYE LAB
Fernanda LOPEZ GARCIA, Intern, DOYE LAB Stephanie MORCHOISNE, Biology engineer, DOYE LAB+33 (0)1 57 27 80 61, room 342B
Stephanie MORCHOISNE, Biology engineer, DOYE LAB+33 (0)1 57 27 80 61, room 342B Gulin OZEK, Intern, DOYE LAB+33 (0)1 57 27 80 62, room 314B
Gulin OZEK, Intern, DOYE LAB+33 (0)1 57 27 80 62, room 314B Ona PRAT PLANAS, PhD student, DOYE LAB+33 (0)1 57 27 80 62, room 342B
Ona PRAT PLANAS, PhD student, DOYE LAB+33 (0)1 57 27 80 62, room 342B Nathalie VADROT, Biology engineer, DOYE LAB+33 (0)1 57 27 80 61, room 342B
Nathalie VADROT, Biology engineer, DOYE LAB+33 (0)1 57 27 80 61, room 342B