Paleogenomics
STEPHANE PEYREGNE

Understanding human biology and health has advanced rapidly thanks to the wealth of genomic, medical, and functional datasets available today. An evolutionary perspective adds a powerful dimension to these efforts by illuminating the processes that shaped our biology and explaining how and why we became who we are. In particular, ancient genomes, including those of Neandertals and Denisovans, provide access to chapters of our evolution that were previously out of reach with present-day data alone. By integrating these ancient genomes with modern datasets, we aim to identify the genetic changes that shaped the modern human lineage and investigate their significance for human biology and health.
Keywords: ancient DNA, human evolution, evolutionary genomics, paleogenomics, population genetics, Neandertals and Denisovans, natural selection, archaic introgression, complex diseases, evolutionary medicine
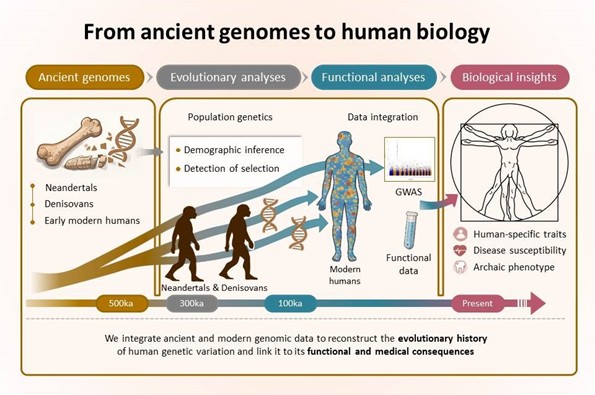
We study how the evolutionary history of our species has shaped human genetic variation and its consequences for biology and health. Genomic approaches have identified many variants associated with traits and diseases, but their interpretation would benefit from being placed within an explicit evolutionary framework. By incorporating this perspective, we aim to understand how and why the biological characteristics of our species have emerged.
Our team relies on ancient genomes to directly access past stages of human evolution that are inaccessible using present-day data alone. These data provide unprecedented temporal resolution to track the emergence, persistence, and disappearance of genetic variants. We are particularly interested in the early periods of our evolution, up to the divergence from archaic humans more than 500,000 years ago, thanks to the analysis of Neandertal and Denisovan genomes.
We generate and analyze paleogenomic data to reconstruct the evolutionary trajectories of genetic variants. We develop methods to exploit these data, from their authentication, including contamination estimation, to evolutionary analyses, such as reconstructing population history and detecting signatures of ancient positive selection. In addition, we use simulations to assess the robustness of these methods and to evaluate different evolutionary scenarios. Together, these approaches allow us to identify genetic changes likely to have important functional effects and to distinguish them from the majority of neutral variation.
We then link these evolutionary trajectories to the present-day effects of genetic variants by integrating functional and medical data. This approach makes it possible, in particular, to identify changes specific to the modern human lineage and to shed light on their contribution to the emergence of traits unique to our lineage. It also provides a framework for investigating the evolutionary origins of susceptibility to complex diseases—such as autoimmune, metabolic, and psychiatric disorders—by distinguishing ancient components of risk, arising from evolutionary trade-offs, from more recent contributions linked to the accumulation of weakly deleterious variants under suboptimal natural selection during our demographic history.
Furthermore, we develop approaches to infer past phenotypic traits, particularly in archaic humans, using ancient genomes and the effects of archaic ancestry observed in present-day populations. These methods provide access to traits not preserved in the fossil record and help place the specific features of our lineage in context. In collaboration with paleoanthropologists and geneticists, we validate these approaches using well-documented traits, such as dental morphology, and apply them for instance to the evolution of the immune system.
By integrating paleogenomic data, evolutionary models, and functional and medical information, our goal is to identify the mechanisms that have shaped human genetic variation and to better understand how our evolutionary history continues to influence our biology and health.
To contact a team member : name.surname@ijm.fr
Yang, J., Iasi, L.N.M., Fu, Q., Cooke, N.P., Kelso, J., Peyrégne, S.*, Peter, B.M.* (2025), An early East Asian lineage with unexpectedly low Denisovan ancestry, Current Biology (* – equally contributing authors).
Swiel, Y., Kelso, J., Peyrégne, S. (2025), Resolving the source of branch length variation in the Y chromosome phylogeny, Genome Biology.
Peyrégne, S., Slon, V., Kelso, J. (2024), More than a decade of genetic research on the Denisovans, Nature Reviews Genetics
Peyrégne, S., Kelso, J., Peter, B.M., Pääbo, S. (2022), The evolutionary history of human spindle genes includes back-and-forth gene flow with Neandertals, eLife.
Peyrégne, S., Slon, V., Mafessoni, F., de Filippo, C., Hajdinjak, M., Nagel, S., Nickel, B., Essel, E., Le Cabec, A., Wehrberger, K., Conard, N.J., Kind, C. J., Posth, C., Krause, J., Abrams, G., Bonjean, D., Di Modica, K., Toussaint, M. Kelso, J., Meyer, M., Pääbo, S., Prüfer, K. (2019), Nuclear DNA from two early Neandertals reveals 80,000 years of genetic continuity in Europe, Science Advances, 5 (6), eaaw5873
Peyrégne, S., Boyle, M., Dannemann, M., Prüfer, K. Detecting ancient positive selection in humans using extended lineage sorting. (2017), Genome Research 27 (9), 1563-157
Massilani, D., Peyrégne, S., Iasi, L. N. M., de Filippo, C., Mafessoni, F., Bossoms Mesa, A., Sümer, A. P., Swiel, Y., Popli, D., Silverman, S., Boyle, M. J., Kozlikin, M. B., Shunkov, M. V., Derevianko, A. P., Higham, T., Douka, K., Meyer, M., Zeberg, H., Kelso, J., & Pääbo, S. (2026). A high-coverage Neandertal genome from the Altai Mountains reveals population structure among Neandertals. Proceedings of the National Academy of Sciences, 123(13), e2534576123. https://doi.org/10.1073/pnas.2534576123
Yang, J., Iasi, L.N.M., Fu, Q., Cooke, N.P., Kelso, J., Peyrégne, S.*, Peter, B.M.* (2025), An early East Asian lineage with unexpectedly low Denisovan ancestry, Current Biology (* – equally contributing authors).
Jacobs, Z., Zavala, E.I., Li, B., O’Gorman, K., Shunkov, M.V., Kozlikin, M.B., Derevianko, A.P., Uliyanov, V., Goldberg, P., Agadjanian, A., Vasiliev, S., Brink, F., Peyrégne, S., Slon, V., Pääbo, S., Kelso, J., Meyer, M., Roberts, R. (2025), Pleistocene chronology and history of hominins and fauna at Denisova Cave, Nature Communications.
Swiel, Y., Kelso, J., Peyrégne, S. (2025), Resolving the source of branch length variation in the Y chromosome phylogeny, Genome Biology.
Sümer, A.P., Rougier, H., Villalba-Mouco, V., Huang, Y., Iasi, L.N.M., Essel, E., Bossoms Mesa, A., Furtwaengler, A., Peyrégne, S., de Filippo, C., Rohrlach, A.B., Pierini, F., Mafessoni, F., Fewlass, H., Elena I. Zavala, E.I., Mylopotamitaki, D., Bianco, R.A., Schmidt, A., Zorn, J., Nickel, B., Patova, A., Posth, C., Smith, G.M., Ruebens, K., Sinet-Mathiot, V., Stoessel, A., Dietl, H., Jörg Orschiedt, J., Kelso, J., Zeberg, H., Bos, K.I., Welker, F., Weiss, M., McPherron, S., Schüler, T., Hublin, J.J., Velemínský, P., Brůžek, J., Peter, B.M., Meyer, M., Meller, H., Ringbauer, H., Hajdinjak, M., Prüfer, K., Krause, J. (2024), Earliest modern human genomes constrain timing of Neandertal admixture, Nature.
Peyrégne, S., Slon, V., Kelso, J. (2024), More than a decade of genetic research on the Denisovans, Nature Reviews Genetics
Popli, D., Peyrégne, S., Peter, B.M. (2023), KIN: A method to infer relatedness from low-coverage ancient DNA, Genome Biology.
Peyrégne, S., Kelso, J., Peter, B.M., Pääbo, S. (2022), The evolutionary history of human spindle genes includes back-and-forth gene flow with Neandertals, eLife.
Skov, L., Peyrégne, S., Popli, D., Iasi, L.N.M., Devièse, T., Slon, V., Zavala, E.I., Hajdinjak, M., Sümer, A., Grote, S., Bossoms Mesa, A., López Herráez, D., Nickel, B., Nagel, S., Richter, J., Essel, E., Gansauge, M., Schmidt, A., Korlević, P., Comeskey, D., Derevianko, A.P., Kharevich, A., Markin, S.V., Talamo, S., Douka, K., Krajcarz, M.T., Roberts, R.G., Higham, T., Viola, B., Krivoshapkin, A.I., Kolobova, K.A., Kelso, J., Meyer, M., Pääbo, S., Peter B.M. (2022), Genetic insights into the social organization of Neanderthals, Nature.
Peyrégne, S., Peter, B.M. (2020), AuthentiCT: a model of ancient DNA damage to estimate the proportion of present-day DNA contamination, Genome Biology.
Khan, N., de Manuel, M., Peyrégne, S., Do, R., Prüfer, K., Marques-Bonet, T., Varki, N., Gagneux, P., & Varki, A. (2020). Multiple Genomic Events Altering Hominin SIGLEC Biology and Innate Immunity Predated the Common Ancestor of Humans and Archaic Hominins. Genome biology and evolution.
Romandini, M., Oxilia, G., Bortolini, E., Peyrégne, S., Delpiano, D., Nava, A., Panetta, D., Di Domenico, G., Martini, P., Arrighi, S., Badino, F., Figus, C., Lugli, F., Marciani, G., Silvestrini, S., Menghi Sartorio, J.C., Terlato, G., Hublin, J.J., Meyer, M., Bondioli, L., Higham, T., Slon, V., Peresani, M., Benazzi, S. (2020), A late Neanderthal tooth from northeastern Italy, Journal of Human Evolution 147, 102867
Peyrégne, S., Prüfer, K. (2020), Present‐Day DNA Contamination in Ancient DNA Datasets, BioEssays, 2000081
Mafessoni, F., Grote, S., de Filippo, C., Slon, V., Kolobova, K. A., Viola, B., Markin, S. V., Chintalapati, M., Peyrégne, S., Skov, L., Skoglund, P., Krivoshapkin, A. I., Derevianko, A. P., Meyer, M., Kelso, J., Peter, B., Prüfer, K., & Pääbo, S. (2020). A high-coverage Neandertal genome from Chagyrskaya Cave. Proceedings of the National Academy of Sciences, 117(26), 15132–15136. https://doi.org/10.1073/pnas.2004944117
Bokelmann, L., Hajdinjak, M., Peyrégne, S., Brace, S., Essel, E., de Filippo, C., Glocke, I., Grote, S., Mafessoni, F., Nagel, S., Kelso, J., Prüfer, K., Vernot, B., Barnes, I., Pääbo, S., Meyer, M., Stringer, C. (2019), A genetic analysis of the Gibraltar Neanderthals, Proceedings of the National Academy of Sciences 116 (31), 15610-15615
Peyrégne, S., Slon, V., Mafessoni, F., de Filippo, C., Hajdinjak, M., Nagel, S., Nickel, B., Essel, E., Le Cabec, A., Wehrberger, K., Conard, N.J., Kind, C. J., Posth, C., Krause, J., Abrams, G., Bonjean, D., Di Modica, K., Toussaint, M. Kelso, J., Meyer, M., Pääbo, S., Prüfer, K. (2019), Nuclear DNA from two early Neandertals reveals 80,000 years of genetic continuity in Europe, Science Advances, 5 (6), eaaw5873
Laval, G., Peyrégne, S., Zidane, N., Harmant, C., Renaud, F., Patin, E., Prugnolle, F., Quintana-Murci, L. (2019), Recent Adaptive Acquisition by African Rainforest Hunter-Gatherers of the Late Pleistocene Sickle-Cell Mutation Suggests Past Differences in Malaria Exposure, The American Journal of Human Genetics 104 (3), 553-561
Slon, V. *, Mafesonni, F.*, Vernot, B. *, de Filippo, C., Grote, S., Viola, B., Hajdinjak, M., Peyrégne, S., Nagel, S., Brown, S., Douka, K., Higham, T., Kozlikin, M.B., Shunkov, M.V., Derevianko, A.P., Kelso, J., Meyer, M., Prüfer K. and Pääbo S. (2018), The genome of the offspring of a Neanderthal mother and a Denisovan father, Nature 561 (7721), 113 (* – equally contributing authors)
Prüfer, K., de Filippo, C. *, Grote, S. *, Mafessoni, F. *, Korlević, P., Hajdinjak, M., Vernot, B., Skov, L., Hsieh, P., Peyrégne, S., Reher, D., Hopfe, C., Nagel, S., Maricic, T., Fu, Q., Theunert, C., Rogers, R., Skoglund, P., Chintalapati, M., Dannemann, M., Nelson, B.J., Key, F.M., Rudan, P., Kućan, Ž., Gušić, I., Golovanova, L.V., Doronichev, V.B., Patterson, N., Reich, D., Eichler, E.E., Slatkin, M., Schierup, M.H., Andrés, A.M., Kelso, J., Meyer, M., Pääbo, S. (2017), A high-coverage Neandertal genome from Vindija Cave in Croatia, Science 358 (6363), 655-658 (* – equally contributing authors)
Peyrégne, S., Boyle, M., Dannemann, M., Prüfer, K. Detecting ancient positive selection in humans using extended lineage sorting. (2017), Genome Research 27 (9), 1563-157
- Diego López-Onaindia (Catalan Institute of Human Paleoecology and Social Evolution)
- Kim Genuite (Université de Bordeaux)
- Lluis Quintana-Murci (Institut Pasteur)
- Marie-Hélène Moncel (Muséum National d’Histoire Naturelle)
- Stéphanie Le Hellard (University of Bergen)
2026-2029 : Fondation pour la Recherche Médicale
2026-2028 : Financement “Crossing Cutting Edges” (Université Paris Cité)
9/01/2026 : Stéphane Peyrégne, team leader at the institute, is the winner of the 2025 Georges Brahms Prize awarded by the CNRS Foundation



